+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Type V-K CAST TnsB bound to LTR-SR | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
| Biological species |  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 11.2 Å | |||||||||||||||
 Authors Authors | Tenjo-Castano F / Sofos N / Lopez-Mendez B / Stutzke LS / Fuglsang A / Stella S / Montoya G | |||||||||||||||
| Funding support |  Denmark, 4 items Denmark, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structure of the TnsB transposase-DNA complex of type V-K CRISPR-associated transposon. Authors: Francisco Tenjo-Castaño / Nicholas Sofos / Blanca López-Méndez / Luisa S Stutzke / Anders Fuglsang / Stefano Stella / Guillermo Montoya /  Abstract: CRISPR-associated transposons (CASTs) are mobile genetic elements that co-opted CRISPR-Cas systems for RNA-guided transposition. Here we present the 2.4 Å cryo-EM structure of the Scytonema ...CRISPR-associated transposons (CASTs) are mobile genetic elements that co-opted CRISPR-Cas systems for RNA-guided transposition. Here we present the 2.4 Å cryo-EM structure of the Scytonema hofmannii (sh) TnsB transposase from Type V-K CAST, bound to the strand transfer DNA. The strand transfer complex displays an intertwined pseudo-symmetrical architecture. Two protomers involved in strand transfer display a catalytically competent active site composed by DDE residues, while other two, which play a key structural role, show active sites where the catalytic residues are not properly positioned for phosphodiester hydrolysis. Transposon end recognition is accomplished by the NTD1/2 helical domains. A singular in trans association of NTD1 domains of the catalytically competent subunits with the inactive DDE domains reinforces the assembly. Collectively, the structural features suggest that catalysis is coupled to protein-DNA assembly to secure proper DNA integration. DNA binding residue mutants reveal that lack of specificity decreases activity, but it could increase transposition in some cases. Our structure sheds light on the strand transfer reaction of DDE transposases and offers new insights into CAST transposition. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15344.map.gz emd_15344.map.gz | 118.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15344-v30.xml emd-15344-v30.xml emd-15344.xml emd-15344.xml | 19.5 KB 19.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_15344.png emd_15344.png | 25.2 KB | ||
| Others |  emd_15344_half_map_1.map.gz emd_15344_half_map_1.map.gz emd_15344_half_map_2.map.gz emd_15344_half_map_2.map.gz | 226.8 MB 226.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15344 http://ftp.pdbj.org/pub/emdb/structures/EMD-15344 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15344 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15344 | HTTPS FTP |
-Validation report
| Summary document |  emd_15344_validation.pdf.gz emd_15344_validation.pdf.gz | 708.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_15344_full_validation.pdf.gz emd_15344_full_validation.pdf.gz | 708.3 KB | Display | |
| Data in XML |  emd_15344_validation.xml.gz emd_15344_validation.xml.gz | 16 KB | Display | |
| Data in CIF |  emd_15344_validation.cif.gz emd_15344_validation.cif.gz | 18.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15344 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15344 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15344 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-15344 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15344.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15344.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.98 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_15344_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
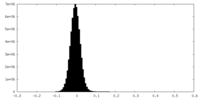
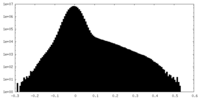
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_15344_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
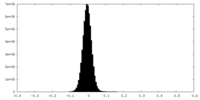
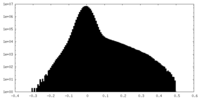
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Type V-K CAST TnsB bound to LTR-SR
| Entire | Name: Type V-K CAST TnsB bound to LTR-SR |
|---|---|
| Components |
|
-Supramolecule #1: Type V-K CAST TnsB bound to LTR-SR
| Supramolecule | Name: Type V-K CAST TnsB bound to LTR-SR / type: complex / Chimera: Yes / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Recombinant expression | Organism:  |
| Molecular weight | Experimental: 359 KDa |
-Macromolecule #1: TnsB
| Macromolecule | Name: TnsB / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Recombinant expression | Organism:  |
| Sequence | String: MNSQQNPDLA VHPLAIPMEG LLGESATTLE KNVIATQLSE EAQVKLEVIQ SLLEPCDRTT YGQKLREAAE KLNVSLRTVQ RLVKNWEQDG LVGLTQTSRA DKGKHRIGEF WEN FITKTY KEGNKGSKRM TPKQVALRVE AKARELKDSK PPNYKTVLRV LAPILEKQQK ...String: MNSQQNPDLA VHPLAIPMEG LLGESATTLE KNVIATQLSE EAQVKLEVIQ SLLEPCDRTT YGQKLREAAE KLNVSLRTVQ RLVKNWEQDG LVGLTQTSRA DKGKHRIGEF WEN FITKTY KEGNKGSKRM TPKQVALRVE AKARELKDSK PPNYKTVLRV LAPILEKQQK AKSIRSPGWR GTTLSVKTRE GKDLSVDYSN HVWQCDHTRV DVLLVDQHGE ILSRPWLT T VIDTYSRCIM GINLGFDAPS SGVVALALRH AILPKRYGSE YKLHCEWGTY GKPEHFYTDG GKDFRSNHLS QIGAQLGFVC HLRDRPSEGG VVERPFKTLN DQLFSTLPGY TGSN VQERP EDAEKDARLT LRELEQLLVR YIVDRYNQSI DARMGDQTRF ERWEAGLPTV PVPIPERDLD ICLMKQSRRT VQRGGCLQFQ NLMYRGEYLA GYAGETVNLR FDPRDITTI LVYRQENNQE VFLTRAHAQG LETEQLALDE AEAASRRLRT AGKTISNQSL LQEVVDRDAL VATKKSRKER QKLEQTVLRS AAVDESNRES LPSQIVEPDE VESTETVHSQ YEDIEVWDYE QLREEYGFGS EFELENLYFQ |
-Macromolecule #2: RightElement
| Macromolecule | Name: RightElement / type: dna / ID: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Sequence | String: TGCTACGTCT CTACGTGTAC AGTGACTAAT TATATGTCGT TGTGACAAAT TATTGTCATC AGTAAAATCC TTATACAGTA TAGATTATAG CGCTTTGGCA GTTTTAGCAT AACCTCTTTG CAGTGACAAA ATAGATGTCG TTGTCCGTGA TTGTGACAAA TTAGCTGTCG ...String: TGCTACGTCT CTACGTGTAC AGTGACTAAT TATATGTCGT TGTGACAAAT TATTGTCATC AGTAAAATCC TTATACAGTA TAGATTATAG CGCTTTGGCA GTTTTAGCAT AACCTCTTTG CAGTGACAAA ATAGATGTCG TTGTCCGTGA TTGTGACAAA TTAGCTGTCG CTTTGCAAGA TAGGAAAAAG CTTTTGTGTA TTTTCATAAT GACAAATTGA CTGTCGCAGG AGGTAA |
-Macromolecule #3: LeftElement
| Macromolecule | Name: LeftElement / type: dna / ID: 3 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Scytonema hofmannii (bacteria) Scytonema hofmannii (bacteria) |
| Sequence | String: ATTATATTGA TGACATTTAA TTTGTCATCA ATTAATTAAG CAACGCTGAT GGGTCACGAC GACAATTAAA TAGTCACAAT GACATTAATC TGTCACCGAC GACAGATAAT TTGTCACTGT ACACTACGCC TTTTGTGG |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS GLACIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Number real images: 738 / Average exposure time: 40.0 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 150000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|
 Movie
Movie Controller
Controller






 Z
Z Y
Y X
X