+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Mouse endoribonuclease Dicer | |||||||||
 Map data Map data | Locally refined electron density map of Dicer helicase domain | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | endoribonuclease / gene silencing / post-transcriptional / catalytic enzyme / cytoplasm / RNA BINDING PROTEIN | |||||||||
| Biological species |  | |||||||||
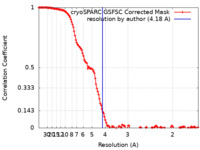
| Method | single particle reconstruction / cryo EM / Resolution: 4.18 Å | |||||||||
 Authors Authors | Zanova M / Zapletal D / Kubicek K / Stefl R / Pinkas M / Novacek J | |||||||||
| Funding support |  Czech Republic, 2 items Czech Republic, 2 items
| |||||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Mammalian Dicer uses its helicase domain to control RNA processing Authors: Zapletal D / Kubicek K / Zanova M / Sebesta M / Pinkas M / Malik R / Joseph DF / Novacek J / Svoboda P / Stefl R | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15299.map.gz emd_15299.map.gz | 200.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15299-v30.xml emd-15299-v30.xml emd-15299.xml emd-15299.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15299_fsc.xml emd_15299_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_15299.png emd_15299.png | 60.4 KB | ||
| Others |  emd_15299_half_map_1.map.gz emd_15299_half_map_1.map.gz emd_15299_half_map_2.map.gz emd_15299_half_map_2.map.gz | 177.1 MB 177.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15299 http://ftp.pdbj.org/pub/emdb/structures/EMD-15299 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15299 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15299 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15299.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15299.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Locally refined electron density map of Dicer helicase domain | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.828 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: half map A
| File | emd_15299_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map B
| File | emd_15299_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Free form of mouse somatic dicer
| Entire | Name: Free form of mouse somatic dicer |
|---|---|
| Components |
|
-Supramolecule #1: Free form of mouse somatic dicer
| Supramolecule | Name: Free form of mouse somatic dicer / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.20 mg/mL | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
Details: The buffer was always prepared fresh | ||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV / Details: Described in STAR methods. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Number real images: 6354 / Average electron dose: 55.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.5 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Endoribonuclease mouse Dicer from AlphaFold database was used as initial model. Initial local fitting into combined map from locally refined protein parts was done using Chimera and then Coot's Real Space Refine Zone. |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 87.8 Target criteria: Ramachandran Plot, Rotamer Analysis, Density Analysis |
-Atomic model buiding 2
| Details | PHENIX Real-space refinement was used for flexible fitting. |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 87.8 / Target criteria: Correlation coefficient |
-Atomic model buiding 3
| Details | ISOLDE was used for flexible fitting with defined torsion restraints |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 87.8 Target criteria: Ramachandran Plot, Rotamer Analysis, Clash Score |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)