+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Lipidic alpha-synuclein fibril - polymorph L2A | |||||||||||||||
 Map data Map data | half map 1 lipidic alpha-synuclein polymorph L2A | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | amyloid fibril / PROTEIN FIBRIL | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / response to desipramine / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process / supramolecular fiber / regulation of synaptic vesicle recycling ...negative regulation of mitochondrial electron transport, NADH to ubiquinone / neutral lipid metabolic process / regulation of acyl-CoA biosynthetic process / negative regulation of dopamine uptake involved in synaptic transmission / negative regulation of norepinephrine uptake / response to desipramine / positive regulation of SNARE complex assembly / positive regulation of hydrogen peroxide catabolic process / supramolecular fiber / regulation of synaptic vesicle recycling / negative regulation of chaperone-mediated autophagy / regulation of reactive oxygen species biosynthetic process / positive regulation of protein localization to cell periphery / mitochondrial membrane organization / negative regulation of exocytosis / regulation of glutamate secretion / dopamine biosynthetic process / positive regulation of neurotransmitter secretion / response to iron(II) ion / regulation of macrophage activation / negative regulation of dopamine metabolic process / negative regulation of platelet-derived growth factor receptor signaling pathway / SNARE complex assembly / negative regulation of thrombin-activated receptor signaling pathway / Lewy body / regulation of locomotion / negative regulation of microtubule polymerization / synaptic vesicle priming / regulation of norepinephrine uptake / transporter regulator activity / protein kinase inhibitor activity / positive regulation of inositol phosphate biosynthetic process / synaptic vesicle transport / dopamine uptake involved in synaptic transmission / regulation of dopamine secretion / positive regulation of receptor recycling / cuprous ion binding / positive regulation of exocytosis / nuclear outer membrane / mitochondrial ATP synthesis coupled electron transport / dynein complex binding / synaptic vesicle exocytosis / response to magnesium ion / positive regulation of endocytosis / negative regulation of serotonin uptake / response to type II interferon / cysteine-type endopeptidase inhibitor activity / kinesin binding / regulation of presynapse assembly / synaptic vesicle endocytosis / alpha-tubulin binding / beta-tubulin binding / phospholipase binding / phospholipid metabolic process / supramolecular fiber organization / behavioral response to cocaine / cellular response to fibroblast growth factor stimulus / cellular response to epinephrine stimulus / inclusion body / Hsp70 protein binding / enzyme inhibitor activity / response to interleukin-1 / axon terminus / cellular response to copper ion / regulation of microtubule cytoskeleton organization / positive regulation of release of sequestered calcium ion into cytosol / SNARE binding / adult locomotory behavior / glutathione metabolic process / protein tetramerization / protein sequestering activity / tubulin binding / phosphoprotein binding / excitatory postsynaptic potential / microglial cell activation / ferrous iron binding / fatty acid metabolic process / phospholipid binding / PKR-mediated signaling / synapse organization / receptor internalization / regulation of long-term neuronal synaptic plasticity / protein destabilization / tau protein binding / enzyme activator activity / positive regulation of inflammatory response / terminal bouton / long-term synaptic potentiation / synaptic vesicle membrane / actin cytoskeleton / growth cone / actin binding / cellular response to oxidative stress / neuron apoptotic process / cell cortex / histone binding / response to lipopolysaccharide / microtubule binding / amyloid fibril formation / chemical synaptic transmission Similarity search - Function | |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
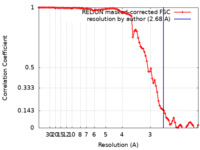
| Method | helical reconstruction / cryo EM / Resolution: 2.68 Å | |||||||||||||||
 Authors Authors | Frieg B / Antonschmidt L / Dienemann C / Geraets JA / Najbauer EE / Matthes D / de Groot BL / Andreas LB / Becker S / Griesinger C / Schroeder GF | |||||||||||||||
| Funding support |  Germany, 4 items Germany, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: The 3D structure of lipidic fibrils of α-synuclein. Authors: Benedikt Frieg / Leif Antonschmidt / Christian Dienemann / James A Geraets / Eszter E Najbauer / Dirk Matthes / Bert L de Groot / Loren B Andreas / Stefan Becker / Christian Griesinger / Gunnar F Schröder /  Abstract: α-synuclein misfolding and aggregation into fibrils is a common feature of α-synucleinopathies, such as Parkinson's disease, in which α-synuclein fibrils are a characteristic hallmark of neuronal ...α-synuclein misfolding and aggregation into fibrils is a common feature of α-synucleinopathies, such as Parkinson's disease, in which α-synuclein fibrils are a characteristic hallmark of neuronal inclusions called Lewy bodies. Studies on the composition of Lewy bodies extracted postmortem from brain tissue of Parkinson's patients revealed that lipids and membranous organelles are also a significant component. Interactions between α-synuclein and lipids have been previously identified as relevant for Parkinson's disease pathology, however molecular insights into their interactions have remained elusive. Here we present cryo-electron microscopy structures of six α-synuclein fibrils in complex with lipids, revealing specific lipid-fibril interactions. We observe that phospholipids promote an alternative protofilament fold, mediate an unusual arrangement of protofilaments, and fill the central cavities of the fibrils. Together with our previous studies, these structures also indicate a mechanism for fibril-induced lipid extraction, which is likely to be involved in the development of α-synucleinopathies. Specifically, one potential mechanism for the cellular toxicity is the disruption of intracellular vesicles mediated by fibrils and oligomers, and therefore the modulation of these interactions may provide a promising strategy for future therapeutic interventions. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_15148.map.gz emd_15148.map.gz | 13.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-15148-v30.xml emd-15148-v30.xml emd-15148.xml emd-15148.xml | 17.8 KB 17.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_15148_fsc.xml emd_15148_fsc.xml | 8.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_15148.png emd_15148.png | 59.8 KB | ||
| Filedesc metadata |  emd-15148.cif.gz emd-15148.cif.gz | 5.8 KB | ||
| Others |  emd_15148_half_map_1.map.gz emd_15148_half_map_1.map.gz emd_15148_half_map_2.map.gz emd_15148_half_map_2.map.gz | 46.2 MB 46.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-15148 http://ftp.pdbj.org/pub/emdb/structures/EMD-15148 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15148 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-15148 | HTTPS FTP |
-Related structure data
| Related structure data |  8a4lMC  8adsC  8aduC  8advC  8adwC  8aexC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_15148.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_15148.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 lipidic alpha-synuclein polymorph L2A | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: lipidic alpha-synuclein polymorph L2A
| File | emd_15148_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | lipidic alpha-synuclein polymorph L2A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2 lipidic alpha-synuclein polymorph L2A
| File | emd_15148_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 lipidic alpha-synuclein polymorph L2A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : amyloid fibril of alpha-synuclein
| Entire | Name: amyloid fibril of alpha-synuclein |
|---|---|
| Components |
|
-Supramolecule #1: amyloid fibril of alpha-synuclein
| Supramolecule | Name: amyloid fibril of alpha-synuclein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Alpha-synuclein
| Macromolecule | Name: Alpha-synuclein / type: protein_or_peptide / ID: 1 / Number of copies: 15 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 14.476108 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDVFMKGLSK AKEGVVAAAE KTKQGVAEAA GKTKEGVLYV GSKTKEGVVH GVATVAEKTK EQVTNVGGAV VTGVTAVAQK TVEGAGSIA AATGFVKKDQ LGKNEEGAPQ EGILEDMPVD PDNEAYEMPS EEGYQDYEPE A UniProtKB: Alpha-synuclein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)