[English] 日本語
 Yorodumi
Yorodumi- EMDB-14955: Cryo-EM structure of Ku 70/80 bound to inositol hexakisphosphate -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Ku 70/80 bound to inositol hexakisphosphate | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | DNA repair / NHEJ / Ku 70/80 / DNA-PK / cancer / double-strand break / DNA binding protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationKu70:Ku80 complex / negative regulation of t-circle formation / DNA end binding / small-subunit processome assembly / positive regulation of lymphocyte differentiation / DNA-dependent protein kinase complex / DNA-dependent protein kinase-DNA ligase 4 complex / nonhomologous end joining complex / regulation of smooth muscle cell proliferation / cellular response to X-ray ...Ku70:Ku80 complex / negative regulation of t-circle formation / DNA end binding / small-subunit processome assembly / positive regulation of lymphocyte differentiation / DNA-dependent protein kinase complex / DNA-dependent protein kinase-DNA ligase 4 complex / nonhomologous end joining complex / regulation of smooth muscle cell proliferation / cellular response to X-ray / double-strand break repair via classical nonhomologous end joining / nuclear telomere cap complex / Cytosolic sensors of pathogen-associated DNA / IRF3-mediated induction of type I IFN / recombinational repair / regulation of telomere maintenance / protein localization to chromosome, telomeric region / U3 snoRNA binding / positive regulation of neurogenesis / cellular hyperosmotic salinity response / 2-LTR circle formation / hematopoietic stem cell proliferation / telomeric repeat DNA binding / DNA 3'-5' helicase / 5'-deoxyribose-5-phosphate lyase activity / hematopoietic stem cell differentiation / ATP-dependent activity, acting on DNA / telomere maintenance via telomerase / site of DNA damage / activation of innate immune response / neurogenesis / telomere maintenance / cyclin binding / DNA-(apurinic or apyrimidinic site) lyase / DNA helicase activity / cellular response to leukemia inhibitory factor / Nonhomologous End-Joining (NHEJ) / small-subunit processome / cellular response to gamma radiation / protein-DNA complex / double-strand break repair via nonhomologous end joining / enzyme activator activity / double-strand break repair / double-stranded DNA binding / scaffold protein binding / DNA recombination / secretory granule lumen / transcription regulator complex / ficolin-1-rich granule lumen / damaged DNA binding / chromosome, telomeric region / transcription cis-regulatory region binding / ribonucleoprotein complex / innate immune response / negative regulation of DNA-templated transcription / DNA damage response / Neutrophil degranulation / ubiquitin protein ligase binding / positive regulation of DNA-templated transcription / protein-containing complex binding / nucleolus / ATP hydrolysis activity / positive regulation of transcription by RNA polymerase II / protein-containing complex / DNA binding / RNA binding / extracellular region / nucleoplasm / ATP binding / membrane / nucleus / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Kefala Stavridi A / Chaplin AK / Blundell TL | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2023 Journal: Nucleic Acids Res / Year: 2023Title: Structural and functional basis of inositol hexaphosphate stimulation of NHEJ through stabilization of Ku-XLF interaction. Authors: Antonia Kefala Stavridi / Amandine Gontier / Vincent Morin / Philippe Frit / Virginie Ropars / Nadia Barboule / Carine Racca / Sagun Jonchhe / Michael J Morten / Jessica Andreani / Alexey ...Authors: Antonia Kefala Stavridi / Amandine Gontier / Vincent Morin / Philippe Frit / Virginie Ropars / Nadia Barboule / Carine Racca / Sagun Jonchhe / Michael J Morten / Jessica Andreani / Alexey Rak / Pierre Legrand / Alexa Bourand-Plantefol / Steven W Hardwick / Dimitri Y Chirgadze / Paul Davey / Taiana Maia De Oliveira / Eli Rothenberg / Sebastien Britton / Patrick Calsou / Tom L Blundell / Paloma F Varela / Amanda K Chaplin / Jean-Baptiste Charbonnier /    Abstract: The classical Non-Homologous End Joining (c-NHEJ) pathway is the predominant process in mammals for repairing endogenous, accidental or programmed DNA Double-Strand Breaks. c-NHEJ is regulated by ...The classical Non-Homologous End Joining (c-NHEJ) pathway is the predominant process in mammals for repairing endogenous, accidental or programmed DNA Double-Strand Breaks. c-NHEJ is regulated by several accessory factors, post-translational modifications, endogenous chemical agents and metabolites. The metabolite inositol-hexaphosphate (IP6) stimulates c-NHEJ by interacting with the Ku70-Ku80 heterodimer (Ku). We report cryo-EM structures of apo- and DNA-bound Ku in complex with IP6, at 3.5 Å and 2.74 Å resolutions respectively, and an X-ray crystallography structure of a Ku in complex with DNA and IP6 at 3.7 Å. The Ku-IP6 interaction is mediated predominantly via salt bridges at the interface of the Ku70 and Ku80 subunits. This interaction is distant from the DNA, DNA-PKcs, APLF and PAXX binding sites and in close proximity to XLF binding site. Biophysical experiments show that IP6 binding increases the thermal stability of Ku by 2°C in a DNA-dependent manner, stabilizes Ku on DNA and enhances XLF affinity for Ku. In cells, selected mutagenesis of the IP6 binding pocket reduces both Ku accrual at damaged sites and XLF enrolment in the NHEJ complex, which translate into a lower end-joining efficiency. Thus, this study defines the molecular bases of the IP6 metabolite stimulatory effect on the c-NHEJ repair activity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14955.map.gz emd_14955.map.gz | 51 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14955-v30.xml emd-14955-v30.xml emd-14955.xml emd-14955.xml | 17 KB 17 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14955_fsc.xml emd_14955_fsc.xml | 13.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_14955.png emd_14955.png | 56.9 KB | ||
| Filedesc metadata |  emd-14955.cif.gz emd-14955.cif.gz | 6.5 KB | ||
| Others |  emd_14955_half_map_1.map.gz emd_14955_half_map_1.map.gz emd_14955_half_map_2.map.gz emd_14955_half_map_2.map.gz | 95.5 MB 95.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14955 http://ftp.pdbj.org/pub/emdb/structures/EMD-14955 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14955 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14955 | HTTPS FTP |
-Related structure data
| Related structure data |  7zt6MC  7z6oC  7zvtC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14955.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14955.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.652 Å | ||||||||||||||||||||||||||||||||||||
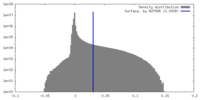
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_14955_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
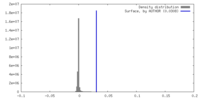
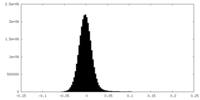
| Density Histograms |
-Half map: #1
| File | emd_14955_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Cryo-EM structure of Ku 70/80 bound to inositol hexakisphosphate
| Entire | Name: Cryo-EM structure of Ku 70/80 bound to inositol hexakisphosphate |
|---|---|
| Components |
|
-Supramolecule #1: Cryo-EM structure of Ku 70/80 bound to inositol hexakisphosphate
| Supramolecule | Name: Cryo-EM structure of Ku 70/80 bound to inositol hexakisphosphate type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: X-ray repair cross-complementing protein 6
| Macromolecule | Name: X-ray repair cross-complementing protein 6 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 73.890336 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNTIHHHHHH NTSGSGGGGG RLVPRGSMSE NLYFQGSMSG WESYYKTEGD EEAEEEQEEN LEASGDYKYS GRDSLIFLVD ASKAMFESQ SEDELTPFDM SIQCIQSVYI SKIISSDRDL LAVVFYGTEK DKNSVNFKNI YVLQELDNPG AKRILELDQF K GQQGQKRF ...String: MNTIHHHHHH NTSGSGGGGG RLVPRGSMSE NLYFQGSMSG WESYYKTEGD EEAEEEQEEN LEASGDYKYS GRDSLIFLVD ASKAMFESQ SEDELTPFDM SIQCIQSVYI SKIISSDRDL LAVVFYGTEK DKNSVNFKNI YVLQELDNPG AKRILELDQF K GQQGQKRF QDMMGHGSDY SLSEVLWVCA NLFSDVQFKM SHKRIMLFTN EDNPHGNDSA KASRARTKAG DLRDTGIFLD LM HLKKPGG FDISLFYRDI ISIAEDEDLR VHFEESSKLE DLLRKVRAKE TRKRALSRLK LKLNKDIVIS VGIYNLVQKA LKP PPIKLY RETNEPVKTK TRTFNTSTGG LLLPSDTKRS QIYGSRQIIL EKEETEELKR FDDPGLMLMG FKPLVLLKKH HYLR PSLFV YPEESLVIGS STLFSALLIK CLEKEVAALC RYTPRRNIPP YFVALVPQEE ELDDQKIQVT PPGFQLVFLP FADDK RKMP FTEKIMATPE QVGKMKAIVE KLRFTYRSDS FENPVLQQHF RNLEALALDL MEPEQAVDLT LPKVEAMNKR LGSLVD EFK ELVYPPDYNP EGKVTKRKHD NEGSGSKRPK VEYSEEELKT HISKGTLGKF TVPMLKEACR AYGLKSGLKK QELLEAL TK HFQD UniProtKB: DNA repair protein Ku70 |
-Macromolecule #2: X-ray repair cross-complementing protein 5
| Macromolecule | Name: X-ray repair cross-complementing protein 5 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on acid anhydrides; Acting on acid anhydrides to facilitate cellular and subcellular movement |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 82.812438 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MVRSGNKAAV VLCMDVGFTM SNSIPGIESP FEQAKKVITM FVQRQVFAEN KDEIALVLFG TDGTDNPLSG GDQYQNITVH RHLMLPDFD LLEDIESKIQ PGSQQADFLD ALIVSMDVIQ HETIGKKFEK RHIEIFTDLS SRFSKSQLDI IIHSLKKCDI S LQFFLPFS ...String: MVRSGNKAAV VLCMDVGFTM SNSIPGIESP FEQAKKVITM FVQRQVFAEN KDEIALVLFG TDGTDNPLSG GDQYQNITVH RHLMLPDFD LLEDIESKIQ PGSQQADFLD ALIVSMDVIQ HETIGKKFEK RHIEIFTDLS SRFSKSQLDI IIHSLKKCDI S LQFFLPFS LGKEDGSGDR GDGPFRLGGH GPSFPLKGIT EQQKEGLEIV KMVMISLEGE DGLDEIYSFS ESLRKLCVFK KI ERHSIHW PCRLTIGSNL SIRIAAYKSI LQERVKKTWT VVDAKTLKKE DIQKETVYCL NDDDETEVLK EDIIQGFRYG SDI VPFSKV DEEQMKYKSE GKCFSVLGFC KSSQVQRRFF MGNQVLKVFA ARDDEAAAVA LSSLIHALDD LDMVAIVRYA YDKR ANPQV GVAFPHIKHN YECLVYVQLP FMEDLRQYMF SSLKNSKKYA PTEAQLNAVD ALIDSMSLAK KDEKTDTLED LFPTT KIPN PRFQRLFQCL LHRALHPREP LPPIQQHIWN MLNPPAEVTT KSQIPLSKIK TLFPLIEAKK KDQVTAQEIF QDNHED GPT AKKLKTEQGG AHFSVSSLAE GSVTSVGSVN PAENFRVLVK QKKASFEEAS NQLINHIEQF LDTNETPYFM KSIDCIR AF REEAIKFSEE QRFNNFLKAL QEKVEIKQLN HFWEIVVQDG ITLITKEEAS GSSVTAEEAK KFLAPKDKPS GDTAAVFE E GGDVDDLLDM I UniProtKB: DNA repair protein Ku80 |
-Macromolecule #3: INOSITOL HEXAKISPHOSPHATE
| Macromolecule | Name: INOSITOL HEXAKISPHOSPHATE / type: ligand / ID: 3 / Number of copies: 1 / Formula: IHP |
|---|---|
| Molecular weight | Theoretical: 660.035 Da |
| Chemical component information |  ChemComp-IHP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 49.43 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.9 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)