[English] 日本語
 Yorodumi
Yorodumi- EMDB-14860: MVV cleaved synaptic complex (CSC) intasome at 4.5 A resolution (... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | MVV cleaved synaptic complex (CSC) intasome at 4.5 A resolution (re-refined) | |||||||||||||||
 Map data Map data | deepEMnhancer map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | Integrase / intasome / MVV / nucleoprotein complex / retrovirus / VIRAL PROTEIN | |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationdUTP diphosphatase / dUTP diphosphatase activity / nucleotide metabolic process / Hydrolases; Acting on peptide bonds (peptidases); Aspartic endopeptidases / ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency ...dUTP diphosphatase / dUTP diphosphatase activity / nucleotide metabolic process / Hydrolases; Acting on peptide bonds (peptidases); Aspartic endopeptidases / ribonuclease H / exoribonuclease H / exoribonuclease H activity / DNA integration / viral genome integration into host DNA / establishment of integrated proviral latency / RNA-directed DNA polymerase / RNA stem-loop binding / RNA-directed DNA polymerase activity / RNA-DNA hybrid ribonuclease activity / Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases / viral capsid / DNA recombination / DNA-directed DNA polymerase / aspartic-type endopeptidase activity / Hydrolases; Acting on ester bonds / DNA-directed DNA polymerase activity / viral translational frameshifting / symbiont entry into host cell / proteolysis / DNA binding / zinc ion binding Similarity search - Function | |||||||||||||||
| Biological species |  Visna/maedi virus EV1 KV1772 / Visna/maedi virus EV1 KV1772 /  Visna-maedi virus / Visna-maedi virus /  Visna lentivirus (strain 1514) Visna lentivirus (strain 1514) | |||||||||||||||
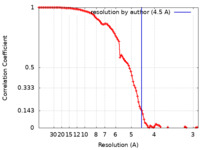
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||||||||
 Authors Authors | Ballandras-Colas A / Maskell D / Pye VE / Locke J / Swuec S / Kotecha A / Costa A / Cherepanov P | |||||||||||||||
| Funding support |  United Kingdom, 4 items United Kingdom, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Science / Year: 2017 Journal: Science / Year: 2017Title: A supramolecular assembly mediates lentiviral DNA integration. Authors: Allison Ballandras-Colas / Daniel P Maskell / Erik Serrao / Julia Locke / Paolo Swuec / Stefán R Jónsson / Abhay Kotecha / Nicola J Cook / Valerie E Pye / Ian A Taylor / Valgerdur ...Authors: Allison Ballandras-Colas / Daniel P Maskell / Erik Serrao / Julia Locke / Paolo Swuec / Stefán R Jónsson / Abhay Kotecha / Nicola J Cook / Valerie E Pye / Ian A Taylor / Valgerdur Andrésdóttir / Alan N Engelman / Alessandro Costa / Peter Cherepanov /    Abstract: Retroviral integrase (IN) functions within the intasome nucleoprotein complex to catalyze insertion of viral DNA into cellular chromatin. Using cryo-electron microscopy, we now visualize the ...Retroviral integrase (IN) functions within the intasome nucleoprotein complex to catalyze insertion of viral DNA into cellular chromatin. Using cryo-electron microscopy, we now visualize the functional maedi-visna lentivirus intasome at 4.9 angstrom resolution. The intasome comprises a homo-hexadecamer of IN with a tetramer-of-tetramers architecture featuring eight structurally distinct types of IN protomers supporting two catalytically competent subunits. The conserved intasomal core, previously observed in simpler retroviral systems, is formed between two IN tetramers, with a pair of C-terminal domains from flanking tetramers completing the synaptic interface. Our results explain how HIV-1 IN, which self-associates into higher-order multimers, can form a functional intasome, reconcile the bulk of early HIV-1 IN biochemical and structural data, and provide a lentiviral platform for design of HIV-1 IN inhibitors. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14860.map.gz emd_14860.map.gz | 92.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14860-v30.xml emd-14860-v30.xml emd-14860.xml emd-14860.xml | 26.9 KB 26.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14860_fsc.xml emd_14860_fsc.xml | 10.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_14860.png emd_14860.png | 175.5 KB | ||
| Masks |  emd_14860_msk_1.map emd_14860_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-14860.cif.gz emd-14860.cif.gz | 7.2 KB | ||
| Others |  emd_14860_additional_1.map.gz emd_14860_additional_1.map.gz emd_14860_additional_2.map.gz emd_14860_additional_2.map.gz emd_14860_half_map_1.map.gz emd_14860_half_map_1.map.gz emd_14860_half_map_2.map.gz emd_14860_half_map_2.map.gz | 51.2 MB 2.8 MB 95.7 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14860 http://ftp.pdbj.org/pub/emdb/structures/EMD-14860 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14860 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14860 | HTTPS FTP |
-Related structure data
| Related structure data |  7zppMC  4139C  5lljC  5m0rC  5t3aC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14860.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14860.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | deepEMnhancer map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.43 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_14860_msk_1.map emd_14860_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: original map - no enhancement
| File | emd_14860_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | original map - no enhancement | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: cryosparc local filter map - to which model...
| File | emd_14860_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | cryosparc local filter map - to which model was fitted and refined against | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_14860_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_14860_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Maedi-visna virus (MVV) intasome
| Entire | Name: Maedi-visna virus (MVV) intasome |
|---|---|
| Components |
|
-Supramolecule #1: Maedi-visna virus (MVV) intasome
| Supramolecule | Name: Maedi-visna virus (MVV) intasome / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Molecular weight | Theoretical: 542.45 KDa |
-Supramolecule #2: Intergrase
| Supramolecule | Name: Intergrase / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Visna/maedi virus EV1 KV1772 Visna/maedi virus EV1 KV1772 |
-Supramolecule #3: vDNA
| Supramolecule | Name: vDNA / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2-#3 |
|---|---|
| Source (natural) | Organism:  Visna-maedi virus Visna-maedi virus |
-Macromolecule #1: Integrase
| Macromolecule | Name: Integrase / type: protein_or_peptide / ID: 1 / Number of copies: 16 / Enantiomer: LEVO EC number: Transferases; Transferring phosphorus-containing groups; Nucleotidyltransferases |
|---|---|
| Source (natural) | Organism:  Visna/maedi virus EV1 KV1772 / Strain: KV1772 Visna/maedi virus EV1 KV1772 / Strain: KV1772 |
| Molecular weight | Theoretical: 32.368826 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: WIENIPLAEE EHNKWHQDAV SLHLEFGIPR TAAEDIVQQC DVCQENKMPS TLRGSNKRGI DHWQVDYTHY EDKIILVWVE TNSGLIYAE RVKGETGQEF RVQTMKWYAM FAPKSLQSDN GPAFVAESTQ LLMKYLGIEH TTGIPWNPQS QALVERTHQT L KNTLEKLI ...String: WIENIPLAEE EHNKWHQDAV SLHLEFGIPR TAAEDIVQQC DVCQENKMPS TLRGSNKRGI DHWQVDYTHY EDKIILVWVE TNSGLIYAE RVKGETGQEF RVQTMKWYAM FAPKSLQSDN GPAFVAESTQ LLMKYLGIEH TTGIPWNPQS QALVERTHQT L KNTLEKLI PMFNAFESAL AGTLITLNIK RKGGLGTSPM DIFIFNKEQQ RIQQQSKSKQ EKIRFCYYRT RKRGHPGEWQ GP TQVLWGG DGAIVVKDRG TDRYLVIANK DVKFIPPPKE IQKE UniProtKB: Gag-Pol polyprotein |
-Macromolecule #2: vDNA, non-transferred strand
| Macromolecule | Name: vDNA, non-transferred strand / type: dna / ID: 2 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Visna lentivirus (strain 1514) Visna lentivirus (strain 1514) |
| Molecular weight | Theoretical: 6.456146 KDa |
| Sequence | String: (DG)(DC)(DT)(DG)(DC)(DG)(DA)(DG)(DA)(DT) (DC)(DC)(DG)(DC)(DT)(DC)(DC)(DG)(DG)(DT) (DG) |
-Macromolecule #3: vDNA, transferred strand
| Macromolecule | Name: vDNA, transferred strand / type: dna / ID: 3 / Number of copies: 2 / Classification: DNA |
|---|---|
| Source (natural) | Organism:  Visna-maedi virus Visna-maedi virus |
| Molecular weight | Theoretical: 5.815762 KDa |
| Sequence | String: (DC)(DA)(DC)(DC)(DG)(DG)(DA)(DG)(DC)(DG) (DG)(DA)(DT)(DC)(DT)(DC)(DG)(DC)(DA) GENBANK: GENBANK: J04359.1 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 6.5 Component:
Details: 1 M NaCl, 3 mM CaCl2 and 25 mM BisTris-HCl, pH 6.5 | ||||||||||||
| Grid | Model: PELCO Ultrathin Carbon with Lacey Carbon / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293.15 K / Instrument: FEI VITROBOT MARK IV Details: To lower salt concentration before plunge-freezing, the grids were blotted for 0.5 s, immediately hydrated with a 4-ul drop of 200 mM NaCl, 3 mM CaCl2 and 25 mM BisTris-HCl pH 6.5 and ...Details: To lower salt concentration before plunge-freezing, the grids were blotted for 0.5 s, immediately hydrated with a 4-ul drop of 200 mM NaCl, 3 mM CaCl2 and 25 mM BisTris-HCl pH 6.5 and blotted again for 2.5 s followed by plunging into liquid ethane.. | ||||||||||||
| Details | A 4 ul drop of freshly prepared intasome in 1 M NaCl, 3 mM CaCl2 and 25 mM BisTris-HCl pH 6.5 was applied onto glow-discharged lacey carbon grids coated with ultrathin carbon (product 01824, Ted Pella). The grids were incubated for 30 s under 100% humidity in a Vitrobot Mark IV (FEI) at 20 oC. To lower salt concentration before plunge-freezing, the grids were blotted for 0.5 s, immediately hydrated with a 4 ul drop of 200 mM NaCl, 3 mM CaCl2, and 25 mM BisTris-HCl pH 6.5 and blotted again for 2.5 s, followed by plunging into liquid ethane. |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 5.0 µm / Nominal defocus min: 3.5 µm |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID:  5m0q Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Details | 5M0Q model was docked to new map (updated relion version and pixel size corrected) in Chimera and refined using phenix.real_space refine and interactively adjusted in coot. |
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 100 / Target criteria: correlation coefficient |
| Output model |  PDB-7zpp: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)