+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tribolium castaneum hexamerin 2 from aggregated sample | |||||||||||||||
 Map data Map data | hexamerin main map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
 Keywords Keywords | hexamerin / tribolium castaneum / storage protein / UNKNOWN FUNCTION | |||||||||||||||
| Biological species |  | |||||||||||||||
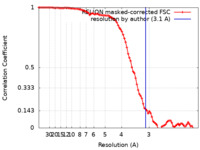
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | |||||||||||||||
 Authors Authors | Valentova L / Plevka P / Fuzik T | |||||||||||||||
| Funding support |  Czech Republic, European Union, 4 items Czech Republic, European Union, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Acta Crystallogr D Struct Biol / Year: 2022 Journal: Acta Crystallogr D Struct Biol / Year: 2022Title: Polyelectrolyte coating of cryo-EM grids improves lateral distribution and prevents aggregation of macromolecules. Authors: Dominik Hrebík / Mária Gondová / Lucie Valentová / Tibor Füzik / Antonín Přidal / Jiří Nováček / Pavel Plevka /  Abstract: Cryo-electron microscopy (cryo-EM) is one of the primary methods used to determine the structures of macromolecules and their complexes. With the increased availability of cryo-electron microscopes, ...Cryo-electron microscopy (cryo-EM) is one of the primary methods used to determine the structures of macromolecules and their complexes. With the increased availability of cryo-electron microscopes, the preparation of high-quality samples has become a bottleneck in the cryo-EM structure-determination pipeline. Macromolecules can be damaged during the purification or preparation of vitrified samples for cryo-EM, making them prone to binding to the grid support, to aggregation or to the adoption of preferential orientations at the air-water interface. Here, it is shown that coating cryo-EM grids with a negatively charged polyelectrolyte, such as single-stranded DNA, before applying the sample reduces the aggregation of macromolecules and improves their distribution. The single-stranded DNA-coated grids enabled the determination of high-resolution structures from samples that aggregated on conventional grids. The polyelectrolyte coating reduces the diffusion of macromolecules and thus may limit the negative effects of the contact of macromolecules with the grid support and blotting paper, as well as of the shear forces on macromolecules during grid blotting. Coating grids with polyelectrolytes can readily be employed in any laboratory dealing with cryo-EM sample preparation, since it is fast, simple, inexpensive and does not require specialized equipment. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14680.map.gz emd_14680.map.gz | 3.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14680-v30.xml emd-14680-v30.xml emd-14680.xml emd-14680.xml | 18.9 KB 18.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_14680_fsc.xml emd_14680_fsc.xml | 9.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_14680.png emd_14680.png | 71.2 KB | ||
| Filedesc metadata |  emd-14680.cif.gz emd-14680.cif.gz | 4.7 KB | ||
| Others |  emd_14680_additional_1.map.gz emd_14680_additional_1.map.gz emd_14680_half_map_1.map.gz emd_14680_half_map_1.map.gz emd_14680_half_map_2.map.gz emd_14680_half_map_2.map.gz | 301.6 KB 50 MB 50.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14680 http://ftp.pdbj.org/pub/emdb/structures/EMD-14680 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14680 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14680 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14680.map.gz / Format: CCP4 / Size: 23 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14680.map.gz / Format: CCP4 / Size: 23 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | hexamerin main map | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||
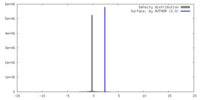
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: mask for half maps
| File | emd_14680_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | mask for half maps | ||||||||||||
| Projections & Slices |
| ||||||||||||
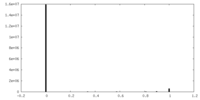
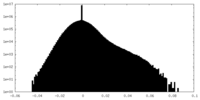
| Density Histograms |
-Half map: #1
| File | emd_14680_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
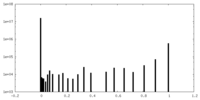
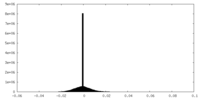
| Density Histograms |
-Half map: #2
| File | emd_14680_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
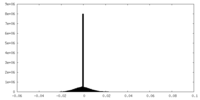
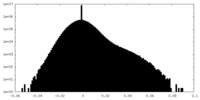
| Density Histograms |
- Sample components
Sample components
-Entire : Hexamerin 2
| Entire | Name: Hexamerin 2 |
|---|---|
| Components |
|
-Supramolecule #1: Hexamerin 2
| Supramolecule | Name: Hexamerin 2 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 / Details: hexamerin isolated from Tribolium castaneum pupae |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 520 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Frames/image: 1-50 / Number grids imaged: 1 / Number real images: 1514 / Average exposure time: 5.0 sec. / Average electron dose: 63.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)