[English] 日本語
 Yorodumi
Yorodumi- EMDB-14398: Cryo-EM structure of the A. baumannii MlaBDEF complex bound to APPNHP -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the A. baumannii MlaBDEF complex bound to APPNHP | |||||||||
 Map data Map data | Cryo-EM map of the A. baumannii MlaBDEF complex | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Acinetobacter baumannii (bacteria) Acinetobacter baumannii (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Mann D / Bergeron JRC | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Commun Biol / Year: 2021 Journal: Commun Biol / Year: 2021Title: Structure and lipid dynamics in the maintenance of lipid asymmetry inner membrane complex of A. baumannii. Authors: Daniel Mann / Junping Fan / Kamolrat Somboon / Daniel P Farrell / Andrew Muenks / Svetomir B Tzokov / Frank DiMaio / Syma Khalid / Samuel I Miller / Julien R C Bergeron /     Abstract: Multi-resistant bacteria are a major threat in modern medicine. The gram-negative coccobacillus Acinetobacter baumannii currently leads the WHO list of pathogens in critical need for new therapeutic ...Multi-resistant bacteria are a major threat in modern medicine. The gram-negative coccobacillus Acinetobacter baumannii currently leads the WHO list of pathogens in critical need for new therapeutic development. The maintenance of lipid asymmetry (MLA) protein complex is one of the core machineries that transport lipids from/to the outer membrane in gram-negative bacteria. It also contributes to broad-range antibiotic resistance in several pathogens, most prominently in A. baumannii. Nonetheless, the molecular details of its role in lipid transport has remained largely elusive. Here, we report the cryo-EM maps of the core MLA complex, MlaBDEF, from the pathogen A. baumannii, in the apo-, ATP- and ADP-bound states, revealing multiple lipid binding sites in the cytosolic and periplasmic side of the complex. Molecular dynamics simulations suggest their potential trajectory across the membrane. Collectively with the recently-reported structures of the E. coli orthologue, this data also allows us to propose a molecular mechanism of lipid transport by the MLA system. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14398.map.gz emd_14398.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14398-v30.xml emd-14398-v30.xml emd-14398.xml emd-14398.xml | 8.2 KB 8.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_14398.png emd_14398.png | 32.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14398 http://ftp.pdbj.org/pub/emdb/structures/EMD-14398 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14398 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14398 | HTTPS FTP |
-Validation report
| Summary document |  emd_14398_validation.pdf.gz emd_14398_validation.pdf.gz | 475 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14398_full_validation.pdf.gz emd_14398_full_validation.pdf.gz | 474.6 KB | Display | |
| Data in XML |  emd_14398_validation.xml.gz emd_14398_validation.xml.gz | 6.2 KB | Display | |
| Data in CIF |  emd_14398_validation.cif.gz emd_14398_validation.cif.gz | 7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14398 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14398 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14398 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14398 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_14398.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14398.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM map of the A. baumannii MlaBDEF complex | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||
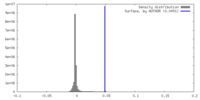
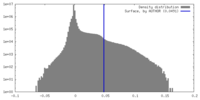
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : MlaBDEF complex
| Entire | Name: MlaBDEF complex |
|---|---|
| Components |
|
-Supramolecule #1: MlaBDEF complex
| Supramolecule | Name: MlaBDEF complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism:  Acinetobacter baumannii (bacteria) Acinetobacter baumannii (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 47.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.9 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 93295 |
|---|---|
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)