[English] 日本語
 Yorodumi
Yorodumi- EMDB-14376: Cryo-EM structure of the Cyanobium sp. PCC 7001 RuBisCO enzyme at... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Cyanobium sp. PCC 7001 RuBisCO enzyme at 3.8 A resolution with C1 symmetry | |||||||||
 Map data Map data | Cryo-EM structure of the Cyanobium sp. PCC 7001 RuBisCO enzyme at 3.8 A resolution with C1 symmetry | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Cyanobium sp. PCC 7001 (bacteria) Cyanobium sp. PCC 7001 (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Evans SL / Mann D / Bergeron JRC | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2023 Journal: Structure / Year: 2023Title: Single-particle cryo-EM analysis of the shell architecture and internal organization of an intact α-carboxysome. Authors: Sasha L Evans / Monsour M J Al-Hazeem / Daniel Mann / Nicolas Smetacek / Andrew J Beavil / Yaqi Sun / Taiyu Chen / Gregory F Dykes / Lu-Ning Liu / Julien R C Bergeron /    Abstract: Carboxysomes are proteinaceous bacterial microcompartments that sequester the key enzymes for carbon fixation in cyanobacteria and some proteobacteria. They consist of a virus-like icosahedral shell, ...Carboxysomes are proteinaceous bacterial microcompartments that sequester the key enzymes for carbon fixation in cyanobacteria and some proteobacteria. They consist of a virus-like icosahedral shell, encapsulating several enzymes, including ribulose 1,5-bisphosphate carboxylase/oxygenase (RuBisCO), responsible for the first step of the Calvin-Benson-Bassham cycle. Despite their significance in carbon fixation and great bioengineering potentials, the structural understanding of native carboxysomes is currently limited to low-resolution studies. Here, we report the characterization of a native α-carboxysome from a marine cyanobacterium by single-particle cryoelectron microscopy (cryo-EM). We have determined the structure of its RuBisCO enzyme, and obtained low-resolution maps of its icosahedral shell, and of its concentric interior organization. Using integrative modeling approaches, we have proposed a complete atomic model of an intact carboxysome, providing insight into its organization and assembly. This is critical for a better understanding of the carbon fixation mechanism and toward repurposing carboxysomes in synthetic biology for biotechnological applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14376.map.gz emd_14376.map.gz | 59.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14376-v30.xml emd-14376-v30.xml emd-14376.xml emd-14376.xml | 9.4 KB 9.4 KB | Display Display |  EMDB header EMDB header |
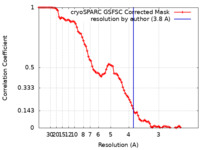
| FSC (resolution estimation) |  emd_14376_fsc.xml emd_14376_fsc.xml | 8.5 KB | Display |  FSC data file FSC data file |
| Images |  emd_14376.png emd_14376.png | 59.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14376 http://ftp.pdbj.org/pub/emdb/structures/EMD-14376 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14376 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14376 | HTTPS FTP |
-Validation report
| Summary document |  emd_14376_validation.pdf.gz emd_14376_validation.pdf.gz | 547.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_14376_full_validation.pdf.gz emd_14376_full_validation.pdf.gz | 546.8 KB | Display | |
| Data in XML |  emd_14376_validation.xml.gz emd_14376_validation.xml.gz | 11 KB | Display | |
| Data in CIF |  emd_14376_validation.cif.gz emd_14376_validation.cif.gz | 14.3 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14376 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14376 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14376 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-14376 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14376.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14376.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-EM structure of the Cyanobium sp. PCC 7001 RuBisCO enzyme at 3.8 A resolution with C1 symmetry | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.11 Å | ||||||||||||||||||||||||||||||||||||
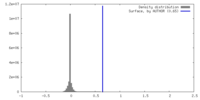
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : RuBisCO
| Entire | Name: RuBisCO |
|---|---|
| Components |
|
-Supramolecule #1: RuBisCO
| Supramolecule | Name: RuBisCO / type: complex / ID: 1 / Chimera: Yes / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Cyanobium sp. PCC 7001 (bacteria) Cyanobium sp. PCC 7001 (bacteria) |
-Macromolecule #1: CbbL
| Macromolecule | Name: CbbL / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Cyanobium sp. PCC 7001 (bacteria) Cyanobium sp. PCC 7001 (bacteria) |
| Sequence | String: MSKKYDAGVK EYRDTYWTPD YVPLDTDLLA CFKCTGQEGV PKEEVAAAVA AESSTGTWST VWSELLVDLD FYKGRCYRIE DVPGDKEAFY AFIAYPLDLF EEGSVTNVLT SLVGNVFGFK ALRHLRLEDI RFPMAFIKTC PGPPNGICVE RDRMNKYGRP LLGCTIKPKL ...String: MSKKYDAGVK EYRDTYWTPD YVPLDTDLLA CFKCTGQEGV PKEEVAAAVA AESSTGTWST VWSELLVDLD FYKGRCYRIE DVPGDKEAFY AFIAYPLDLF EEGSVTNVLT SLVGNVFGFK ALRHLRLEDI RFPMAFIKTC PGPPNGICVE RDRMNKYGRP LLGCTIKPKL GLSGKNYGRV VYECLRGGLD FTKDDENINS QPFQRWQNRF EFVAEAVALA QQETGEKKGH YLNCTAATPE EMYERAEFAK ELGQPIIMHD YITGGFTANT GLSKWCRKNG MLLHIHRAMH AVIDRHPKHG IHFRVLAKCL RLSGGDQLHT GTVVGKLEGD RQTTLGFIDQ LRESFIPEDR SRGNFFDQDW GSMPGVFAVA SGGIHVWHMP ALVAIFGDDS VLQFGGGTHG HPWGSAAGAA ANRVALEACV KARNAGREIE KESRDILMEA AKHSPELAIA LETWKEIKFE FDTVDKLDVQ |
-Macromolecule #2: CbbS
| Macromolecule | Name: CbbS / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Cyanobium sp. PCC 7001 (bacteria) Cyanobium sp. PCC 7001 (bacteria) |
| Sequence | String: MPFKSTVGDY QTVATLETFG FLPPMTQDEI YDQIAYIIAQ GWSPLIEHVH PSRSMATYWS YWKLPFFGEK DLGVIVSELE ACHRAYPDHH VRLVGYDAYT QSQGACFVVF EGR |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.5 µm / Nominal defocus min: 0.5 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)