+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of mammalian AAP in complex with Meropenem | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | inhibitor / complex / acylaminoacyl-peptidase / meropenem / carbapenem / acylpeptide hydrolase / HYDROLASE | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationacylaminoacyl-peptidase / omega peptidase activity / serine-type endopeptidase activity / proteolysis / cytoplasm Similarity search - Function | ||||||||||||||||||
| Biological species |  | ||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.1 Å | ||||||||||||||||||
 Authors Authors | Kiss-Szeman AJ / Harmat V | ||||||||||||||||||
| Funding support |  Hungary, 5 items Hungary, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Chem Sci / Year: 2022 Journal: Chem Sci / Year: 2022Title: A carbapenem antibiotic inhibiting a mammalian serine protease: structure of the acylaminoacyl peptidase-meropenem complex. Authors: Anna J Kiss-Szemán / Luca Takács / Zoltán Orgován / Pál Stráner / Imre Jákli / Gitta Schlosser / Simonas Masiulis / Veronika Harmat / Dóra K Menyhárd / András Perczel /   Abstract: The structure of porcine AAP (pAAP) in a covalently bound complex with meropenem was determined by cryo-EM to 2.1 Å resolution, showing the mammalian serine-protease inhibited by a carbapenem ...The structure of porcine AAP (pAAP) in a covalently bound complex with meropenem was determined by cryo-EM to 2.1 Å resolution, showing the mammalian serine-protease inhibited by a carbapenem antibiotic. AAP is a modulator of the ubiquitin-proteasome degradation system and the site of a drug-drug interaction between the widely used antipsychotic, valproate and carbapenems. The active form of pAAP - a toroidal tetramer - binds four meropenem molecules covalently linked to the catalytic Ser587 of the serine-protease triad, in an acyl-enzyme state. AAP is hindered from fully processing the antibiotic by the displacement and protonation of His707 of the catalytic triad. We show that AAP is made susceptible to the association by its unusually sheltered active pockets and flexible catalytic triads, while the carbapenems possess sufficiently small substituents on their β-lactam rings to fit into the shallow substrate-specificity pocket of the enzyme. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14149.map.gz emd_14149.map.gz | 168.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14149-v30.xml emd-14149-v30.xml emd-14149.xml emd-14149.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_14149.png emd_14149.png | 214 KB | ||
| Filedesc metadata |  emd-14149.cif.gz emd-14149.cif.gz | 6.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14149 http://ftp.pdbj.org/pub/emdb/structures/EMD-14149 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14149 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14149 | HTTPS FTP |
-Related structure data
| Related structure data |  7qunMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14149.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14149.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.96 Å | ||||||||||||||||||||||||||||||||||||
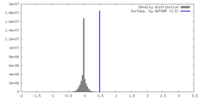
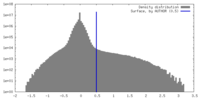
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
-Entire : Homotetramer of aclyaminoacyl-peptidase
| Entire | Name: Homotetramer of aclyaminoacyl-peptidase |
|---|---|
| Components |
|
-Supramolecule #1: Homotetramer of aclyaminoacyl-peptidase
| Supramolecule | Name: Homotetramer of aclyaminoacyl-peptidase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Acylamino-acid-releasing enzyme
| Macromolecule | Name: Acylamino-acid-releasing enzyme / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 81.324391 KDa |
| Sequence | String: MERQVLLSEP EEAAALYRGL SRQPALSAAC LGPEVTTQYG GRYRTVHTEW TQRDLERMEN IRFCRQYLVF HDGDSVVFAG PAGNSVETR GELLSRESPS GTMKAVLRKA GGTGTAEEKQ FLEVWEKNRK LKSFNLSALE KHGPVYEDDC FGCLSWSHSE T HLLYVADK ...String: MERQVLLSEP EEAAALYRGL SRQPALSAAC LGPEVTTQYG GRYRTVHTEW TQRDLERMEN IRFCRQYLVF HDGDSVVFAG PAGNSVETR GELLSRESPS GTMKAVLRKA GGTGTAEEKQ FLEVWEKNRK LKSFNLSALE KHGPVYEDDC FGCLSWSHSE T HLLYVADK KRPKAESFFQ TKALDVTGSD DEMARTKKPD QAIKGDQFLF YEDWGENMVS KSTPVLCVLD IESGNISVLE GV PESVSPG QAFWAPGDTG VVFVGWWHEP FRLGIRFCTN RRSALYYVDL TGGKCELLSD ESVAVTSPRL SPDQCRIVYL RFP SLVPHQ QCGQLCLYDW YTRVTSVVVD IVPRQLGEDF SGIYCSLLPL GCWSADSQRV VFDSPQRSRQ DLFAVDTQMG SVTS LTAGG SGGSWKLLTI DRDLMVVQFS TPSVPPSLKV GFLPPAGKEQ AVSWVSLEEA EPFPDISWSI RVLQPPPQQE HVQYA GLDF EAILLQPSNS PEKTQVPMVV MPHGGPHSSF VTAWMLFPAM LCKMGFAVLL VNYRGSTGFG QDSILSLPGN VGHQDV KDV QFAVEQVLQE EHFDAGRVAL MGGSHGGFLS CHLIGQYPET YSACVVRNPV INIASMMGST DIPDWCMVEA GFSYSSD CL PDLSVWAAML DKSPIKYAPQ VKTPLLLMLG QEDRRVPFKQ GMEYYRVLKA RNVPVRLLLY PKSTHALSEV EVESDSFM N AVLWLCTHLG S UniProtKB: Acylamino-acid-releasing enzyme |
-Macromolecule #2: (2S,3R,4S)-4-{[(3S,5S)-5-(dimethylcarbamoyl)pyrrolidin-3-yl]sulfa...
| Macromolecule | Name: (2S,3R,4S)-4-{[(3S,5S)-5-(dimethylcarbamoyl)pyrrolidin-3-yl]sulfanyl}-2-[(2S,3R)-3-hydroxy-1-oxobutan-2-yl]-3-methyl-3,4-dihydro-2H-pyrrole-5-carboxylic acid type: ligand / ID: 2 / Number of copies: 4 / Formula: DWZ |
|---|---|
| Molecular weight | Theoretical: 385.478 Da |
| Chemical component information | 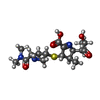 ChemComp-DWZ: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Component - Concentration: 10.0 mM / Component - Formula: C4H11NO3 / Component - Name: TRIS |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 25 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
| Details | Covalent complex with carbapenem type antibiotic Meropenem |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Number grids imaged: 4 / Number real images: 11625 / Average exposure time: 5.93 sec. / Average electron dose: 41.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 130000 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)