[English] 日本語
 Yorodumi
Yorodumi- EMDB-13960: Cryo-EM structure of Ldh-EtfAB complex from Acetobacterium woodii -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Ldh-EtfAB complex from Acetobacterium woodii | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Electron bifurcation / Electron confirmation / Lactate / Lactate dehydrogenase complex / Electron transferring flavoprotein / A. woodii / redox enzyme / FLAVOPROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationlactate dehydrogenase (NAD+,ferredoxin) / fatty acid beta-oxidation using acyl-CoA dehydrogenase / FAD binding / flavin adenine dinucleotide binding / 4 iron, 4 sulfur cluster binding / electron transfer activity / oxidoreductase activity / metal ion binding / cytoplasm Similarity search - Function | |||||||||
| Biological species |  Acetobacterium woodii (bacteria) Acetobacterium woodii (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.43 Å | |||||||||
 Authors Authors | Kayastha K / Ermler U | |||||||||
| Funding support |  Germany, 1 items Germany, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2022 Journal: Elife / Year: 2022Title: Structure-based electron-confurcation mechanism of the Ldh-EtfAB complex. Authors: Kanwal Kayastha / Alexander Katsyv / Christina Himmrich / Sonja Welsch / Jan M Schuller / Ulrich Ermler / Volker Müller /  Abstract: Lactate oxidation with NAD as electron acceptor is a highly endergonic reaction. Some anaerobic bacteria overcome the energetic hurdle by flavin-based electron bifurcation/confurcation (FBEB/FBEC) ...Lactate oxidation with NAD as electron acceptor is a highly endergonic reaction. Some anaerobic bacteria overcome the energetic hurdle by flavin-based electron bifurcation/confurcation (FBEB/FBEC) using a lactate dehydrogenase (Ldh) in concert with the electron-transferring proteins EtfA and EtfB. The electron cryo-microscopically characterized (Ldh-EtfAB) complex of at 2.43 Å resolution consists of a mobile EtfAB shuttle domain located between the rigid central Ldh and the peripheral EtfAB base units. The FADs of Ldh and the EtfAB shuttle domain contact each other thereby forming the D (dehydrogenation-connected) state. The intermediary Glu37 and Glu139 may harmonize the redox potentials between the FADs and the pyruvate/lactate pair crucial for FBEC. By integrating Alphafold2 calculations a plausible novel B (bifurcation-connected) state was obtained allowing electron transfer between the EtfAB base and shuttle FADs. Kinetic analysis of enzyme variants suggests a correlation between NAD binding site and D-to-B-state transition implicating a 75° rotation of the EtfAB shuttle domain. The FBEC inactivity when truncating the ferredoxin domain of EtfA substantiates its role as redox relay. Lactate oxidation in Ldh is assisted by the catalytic base His423 and a metal center. On this basis, a comprehensive catalytic mechanism of the FBEC process was proposed. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_13960.map.gz emd_13960.map.gz | 69.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-13960-v30.xml emd-13960-v30.xml emd-13960.xml emd-13960.xml | 22.2 KB 22.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_13960_fsc.xml emd_13960_fsc.xml | 13.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_13960.png emd_13960.png | 127.6 KB | ||
| Masks |  emd_13960_msk_1.map emd_13960_msk_1.map | 91.1 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-13960.cif.gz emd-13960.cif.gz | 7.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13960 http://ftp.pdbj.org/pub/emdb/structures/EMD-13960 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13960 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13960 | HTTPS FTP |
-Related structure data
| Related structure data |  7qh2MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_13960.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_13960.map.gz / Format: CCP4 / Size: 91.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.93 Å | ||||||||||||||||||||||||||||||||||||
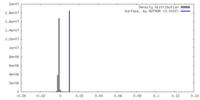
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_13960_msk_1.map emd_13960_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
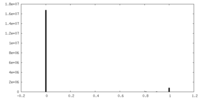
| Projections & Slices |
| ||||||||||||
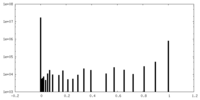
| Density Histograms |
- Sample components
Sample components
-Entire : Ldh-EtfAB complex
| Entire | Name: Ldh-EtfAB complex |
|---|---|
| Components |
|
-Supramolecule #1: Ldh-EtfAB complex
| Supramolecule | Name: Ldh-EtfAB complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 Details: Complex composed of a dimer of a trimer of EtfA, EtfB and Ldh subunits making a hexameric oligomer state. Each of the subunits contain 1 FAD co-factor making total of 6 FADs. 2 Ldh subunits ...Details: Complex composed of a dimer of a trimer of EtfA, EtfB and Ldh subunits making a hexameric oligomer state. Each of the subunits contain 1 FAD co-factor making total of 6 FADs. 2 Ldh subunits contain 1 Fe molecule each. |
|---|---|
| Source (natural) | Organism:  Acetobacterium woodii (bacteria) Acetobacterium woodii (bacteria) |
| Molecular weight | Theoretical: 220 KDa |
-Macromolecule #1: Lactate dehydrogenase (NAD(+),ferredoxin) subunit LctC
| Macromolecule | Name: Lactate dehydrogenase (NAD(+),ferredoxin) subunit LctC type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: ec: 1.3.1.110 |
|---|---|
| Source (natural) | Organism:  Acetobacterium woodii (bacteria) / Strain: ATCC 29683 / DSM 1030 / JCM 2381 / KCTC 1655 / WB1 Acetobacterium woodii (bacteria) / Strain: ATCC 29683 / DSM 1030 / JCM 2381 / KCTC 1655 / WB1 |
| Molecular weight | Theoretical: 46.222496 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAGIKIIKEN VDRETFEALA EICPFDAFSY ENDKLEVTAA CKMCKMCLKK GPEGVLILEE DEKVAIDKSL YRGITVYVDH IEGQIHPVT FELIGKAREL AAVIGHPVYA LLMGTNITEK ADELLKYGVD KVFVYDKPEL KHFVIEPYAN VLEDFIEKVK P SSILVGAT ...String: MAGIKIIKEN VDRETFEALA EICPFDAFSY ENDKLEVTAA CKMCKMCLKK GPEGVLILEE DEKVAIDKSL YRGITVYVDH IEGQIHPVT FELIGKAREL AAVIGHPVYA LLMGTNITEK ADELLKYGVD KVFVYDKPEL KHFVIEPYAN VLEDFIEKVK P SSILVGAT NVGRSLAPRV AARYRTGLTA DCTILEMKEN TDLVQIRPAF GGNIMAQIVT ENTRPQFCTV RYKVFTAPER VN EPWGDVE MMDIEKAKLV SAIEVMEVIK KEKGIDLSEA ETIVAVGRGV KCEKDLDMIH EFAEKIGATV ACTRPGIEAG WFD ARLQIG LSGRTVKPKL IIALGISGAV QFAAGMQNSE YIIAINSDPK APIFNIAHCG MVGDLYEILP ELLTMIEGPE NNKD TETIS IPEAIETPER MVV UniProtKB: Lactate dehydrogenase (NAD(+),ferredoxin) subunit LctC |
-Macromolecule #2: Lactate dehydrogenase (NAD(+),ferredoxin) subunit LctB
| Macromolecule | Name: Lactate dehydrogenase (NAD(+),ferredoxin) subunit LctB type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO / EC number: ec: 1.3.1.110 |
|---|---|
| Source (natural) | Organism:  Acetobacterium woodii (bacteria) / Strain: ATCC 29683 / DSM 1030 / JCM 2381 / KCTC 1655 / WB1 Acetobacterium woodii (bacteria) / Strain: ATCC 29683 / DSM 1030 / JCM 2381 / KCTC 1655 / WB1 |
| Molecular weight | Theoretical: 29.187627 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSKILVCIKQ VPGTSNVEVD PETGVLIRDG VESKLNPYDL FGLETAFRLK EQLGGTITTL SMGPMQSKEV LMESFYMGAD EGCLLSDRK FGGADVVATS YTLAQGTKRL GDFDLIICGK QTTDGDTAQV GPEMAEFLGI PHVTNVIKIL AADEKGLTLQ M NMEESLEI ...String: MSKILVCIKQ VPGTSNVEVD PETGVLIRDG VESKLNPYDL FGLETAFRLK EQLGGTITTL SMGPMQSKEV LMESFYMGAD EGCLLSDRK FGGADVVATS YTLAQGTKRL GDFDLIICGK QTTDGDTAQV GPEMAEFLGI PHVTNVIKIL AADEKGLTLQ M NMEESLEI QRVPYPCLIT VDKDIYTPRL PSYKRKLDIS KNPEIKILTL KDMYDTNEKK YGLSGSPTQV ERIFPPESNV EK TSFEGDG KVLAKALLGI LTEKKYLG UniProtKB: Lactate dehydrogenase (NAD(+),ferredoxin) subunit LctB |
-Macromolecule #3: Lactate dehydrogenase (NAD(+),ferredoxin) subunit LctD
| Macromolecule | Name: Lactate dehydrogenase (NAD(+),ferredoxin) subunit LctD type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO / EC number: ec: 1.3.1.110 |
|---|---|
| Source (natural) | Organism:  Acetobacterium woodii (bacteria) / Strain: ATCC 29683 / DSM 1030 / JCM 2381 / KCTC 1655 / WB1 Acetobacterium woodii (bacteria) / Strain: ATCC 29683 / DSM 1030 / JCM 2381 / KCTC 1655 / WB1 |
| Molecular weight | Theoretical: 51.267797 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MNYKKVEASD IAAIKELIPA ERVFVGTEIG EDFSHDELGS IHSYPEVLIK VTSTEEVSKI MKYAYEHNIP VVVRGSGTGL VGACVPLFG GIMLETTLMN NILELDTENL TVTVEPGVLL MELSKFVEEN DLFYPPDPGE KSATIAGNIS TNAGGMRAVK Y GVTRDYVR ...String: MNYKKVEASD IAAIKELIPA ERVFVGTEIG EDFSHDELGS IHSYPEVLIK VTSTEEVSKI MKYAYEHNIP VVVRGSGTGL VGACVPLFG GIMLETTLMN NILELDTENL TVTVEPGVLL MELSKFVEEN DLFYPPDPGE KSATIAGNIS TNAGGMRAVK Y GVTRDYVR GLTVVLANGE IIELGGKIVK NSSGYSLKDL VIGSEGTLCV ITKAILKLLP LPKMTLSLLI PFENISDAAG IV PKIIKSK AIPTAIEFME RQTILFAEDF LGKKFPDSSS NAYILLTFDG NTKEQVEAEY ETVANLCLAE GAKDVYIVDT VER KDSVWS ARGAFLEAIK ASTTEMDECD VVVPRNRIAE FIEFTHDLAK EMDVRIPSFG HAGDGNLHIY VCRDELCQAD WEAK LAEAM DRMYAKALTF EGLVSGEHGI GYAKRKYLLN DFGTEHLALM AGIKQTFDPK NLLNPKKVCQ MA UniProtKB: Lactate dehydrogenase (NAD(+),ferredoxin) subunit LctD |
-Macromolecule #4: FLAVIN-ADENINE DINUCLEOTIDE
| Macromolecule | Name: FLAVIN-ADENINE DINUCLEOTIDE / type: ligand / ID: 4 / Number of copies: 6 / Formula: FAD |
|---|---|
| Molecular weight | Theoretical: 785.55 Da |
| Chemical component information |  ChemComp-FAD: |
-Macromolecule #5: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 2 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.8 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
Details: pH 7.5 | |||||||||
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 135 sec. / Pretreatment - Atmosphere: NITROGEN / Pretreatment - Pressure: 38.0 kPa / Details: 45mA | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV Details: Blot force of +20 and blotting time of 4 seconds before plunging. | |||||||||
| Details | Sample was cross linked with 1 mM of BS3 before grid preparation. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 30 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Number real images: 9788 / Average exposure time: 6.52 sec. / Average electron dose: 106.17 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.1 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-7qh2: |
 Movie
Movie Controller
Controller






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)