[English] 日本語
 Yorodumi
Yorodumi- EMDB-1319: Octomeric pyruvate-ferredoxin oxidoreductase from Desulfovibrio v... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1319 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Octomeric pyruvate-ferredoxin oxidoreductase from Desulfovibrio vulgaris. | |||||||||
 Map data Map data | pyrovat-ferredoxin oxidoreductase from Desulfovibrio vulgaris Hildenborough | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Desulfovibrio vulgaris str. Hildenborough (bacteria) Desulfovibrio vulgaris str. Hildenborough (bacteria) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 17.0 Å | |||||||||
 Authors Authors | Garczarek F / Dong M / Typke D / Witkowska E / Hazen TC / Nogales E / Glaeser R | |||||||||
 Citation Citation |  Journal: J Struct Biol / Year: 2007 Journal: J Struct Biol / Year: 2007Title: Octomeric pyruvate-ferredoxin oxidoreductase from Desulfovibrio vulgaris. Authors: Florian Garczarek / Ming Dong / Dieter Typke / H Ewa Witkowska / Terry C Hazen / Eva Nogales / Mark D Biggin / Robert M Glaeser /  Abstract: Pyruvate-ferredoxin oxidoreductatse (PFOR) carries out the central step in oxidative decarboxylation of pyruvate to acetyl-CoA. We have purified this enzyme from Desulfovibrio vulgaris Hildenborough ...Pyruvate-ferredoxin oxidoreductatse (PFOR) carries out the central step in oxidative decarboxylation of pyruvate to acetyl-CoA. We have purified this enzyme from Desulfovibrio vulgaris Hildenborough (DvH) as part of a systematic characterization of as many multiprotein complexes as possible for this organism, and the three-dimensional structure of this enzyme has been determined by a combination of electron microscopy (EM), single particle image analysis, homology modeling and computational molecular docking. Our results show that the 1MDa DvH PFOR complex is a homo-octomer, or more precisely, a tetramer of the dimeric form of the related enzyme found in Desulfovibrio africanus (Da), with which it shares a sequence identity of 69%. Our homology model of the DvH PFOR dimer is based on the Da PFOR X-ray structure. Docking of this model into our 17A resolution EM-reconstruction of negatively stained DvH PFOR octomers strongly suggests that the difference in oligomerization state for the two species is due to the insertion of a single valine residue (Val383) within a surface loop of the DvH enzyme. This study demonstrates that the strategy of intermediate resolution EM reconstruction coupled to homology modeling and docking can be powerful enough to infer the functionality of single amino acid residues. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1319.map.gz emd_1319.map.gz | 14 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1319-v30.xml emd-1319-v30.xml emd-1319.xml emd-1319.xml | 8.6 KB 8.6 KB | Display Display |  EMDB header EMDB header |
| Images |  1319.gif 1319.gif | 50.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1319 http://ftp.pdbj.org/pub/emdb/structures/EMD-1319 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1319 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1319 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1319.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1319.map.gz / Format: CCP4 / Size: 15.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | pyrovat-ferredoxin oxidoreductase from Desulfovibrio vulgaris Hildenborough | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.56 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : pyruvate-ferredoxin oxidoreductase from Desulfovibrio vulgaris Hi...
| Entire | Name: pyruvate-ferredoxin oxidoreductase from Desulfovibrio vulgaris Hildenborough |
|---|---|
| Components |
|
-Supramolecule #1000: pyruvate-ferredoxin oxidoreductase from Desulfovibrio vulgaris Hi...
| Supramolecule | Name: pyruvate-ferredoxin oxidoreductase from Desulfovibrio vulgaris Hildenborough type: sample / ID: 1000 / Oligomeric state: homooctomer / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 1.056 MDa |
-Macromolecule #1: pyruvate-ferredoxin oxidoreductase
| Macromolecule | Name: pyruvate-ferredoxin oxidoreductase / type: protein_or_peptide / ID: 1 / Name.synonym: PFOR / Number of copies: 8 / Oligomeric state: homooctomer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Desulfovibrio vulgaris str. Hildenborough (bacteria) Desulfovibrio vulgaris str. Hildenborough (bacteria)Strain: ATCC 29579 |
| Molecular weight | Experimental: 132 MDa / Theoretical: 132 MDa |
| Recombinant expression | Organism:  Desulfovibrio vulgaris str. Hildenborough (bacteria) Desulfovibrio vulgaris str. Hildenborough (bacteria) |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.05 mg/mL |
|---|---|
| Buffer | pH: 7.6 / Details: 10mM HEPES, 2mM DTT, 0.01% NP40 |
| Staining | Type: NEGATIVE / Details: 5% ammonium molybdate, 1% trehalose, pH7 |
| Grid | Details: 400 mesh copper grid coated with carbon layer |
| Vitrification | Cryogen name: NONE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: OTHER / Digitization - Sampling interval: 12.7 µm / Number real images: 19 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Calibrated magnification: 49700 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 6.2 mm / Nominal defocus max: 1.6 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: side entry / Specimen holder model: OTHER |
- Image processing
Image processing
| CTF correction | Details: defocus groups |
|---|---|
| Final reconstruction | Applied symmetry - Point group: D4 (2x4 fold dihedral) / Resolution.type: BY AUTHOR / Resolution: 17.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: EMAN, SPIDER / Number images used: 12402 |
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera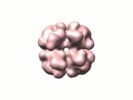






 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















