[English] 日本語
 Yorodumi
Yorodumi- EMDB-1272: Cryoelectron microscopy of protein IX-modified adenoviruses sugge... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1272 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryoelectron microscopy of protein IX-modified adenoviruses suggests a new position for the C terminus of protein IX. | |||||||||
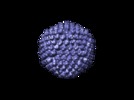
 Map data Map data | This is the reconstruction of Ad-IX-GFP capsids. The data were collected at 2.17 angstrom per pixel and downsampled by 2 for processing. The final reconstruction was guassian lowpass filtered to 22 angstroms. The map deposited here is oriented with the 3-fold axis along z. This map is a half-map. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Human adenovirus 5 Human adenovirus 5 | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 22.0 Å | |||||||||
 Authors Authors | Marsh MP / Campos SK / Baker ML / Chen CY / Chiu W / Barry MA | |||||||||
 Citation Citation |  Journal: J Virol / Year: 2006 Journal: J Virol / Year: 2006Title: Cryoelectron microscopy of protein IX-modified adenoviruses suggests a new position for the C terminus of protein IX. Authors: Michael P Marsh / Samuel K Campos / Matthew L Baker / Christopher Y Chen / Wah Chiu / Michael A Barry /  Abstract: Recombinant human adenovirus is a useful gene delivery vector for clinical gene therapy. Minor capsid protein IX of adenovirus has been of recent interest since multiple studies have shown that ...Recombinant human adenovirus is a useful gene delivery vector for clinical gene therapy. Minor capsid protein IX of adenovirus has been of recent interest since multiple studies have shown that modifications can be made to its C terminus to alter viral tropism or add molecular tags and/or reporter proteins. We examined the structure of an engineered adenovirus displaying the enhanced green fluorescent protein (EGFP) fused to the C terminus of protein IX. Cryoelectron microscopy and reconstruction localized the C-terminal EGFP fusion between the H2 hexon and the H4 hexon, positioned between adjacent facets, directly above the density previously assigned as protein IIIa. The original assignment of IIIa was based largely on indirect evidence, and the data presented herein support the reassignment of the IIIa density as protein IX. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1272.map.gz emd_1272.map.gz | 32.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1272-v30.xml emd-1272-v30.xml emd-1272.xml emd-1272.xml | 9.3 KB 9.3 KB | Display Display |  EMDB header EMDB header |
| Images |  1272.gif 1272.gif | 27.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1272 http://ftp.pdbj.org/pub/emdb/structures/EMD-1272 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1272 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1272 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1272.map.gz / Format: CCP4 / Size: 60.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1272.map.gz / Format: CCP4 / Size: 60.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is the reconstruction of Ad-IX-GFP capsids. The data were collected at 2.17 angstrom per pixel and downsampled by 2 for processing. The final reconstruction was guassian lowpass filtered to 22 angstroms. The map deposited here is oriented with the 3-fold axis along z. This map is a half-map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Adenovirus capsid with GFP fused to C-terminus of capisd protein pIX
| Entire | Name: Adenovirus capsid with GFP fused to C-terminus of capisd protein pIX |
|---|---|
| Components |
|
-Supramolecule #1000: Adenovirus capsid with GFP fused to C-terminus of capisd protein pIX
| Supramolecule | Name: Adenovirus capsid with GFP fused to C-terminus of capisd protein pIX type: sample / ID: 1000 / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 150 MDa |
-Supramolecule #1: Human adenovirus 5
| Supramolecule | Name: Human adenovirus 5 / type: virus / ID: 1 / Name.synonym: Adenovirus, Ad-IX-GFP / Details: GFP fusion to protein pIX / NCBI-ID: 28285 / Sci species name: Human adenovirus 5 / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No / Syn species name: Adenovirus, Ad-IX-GFP |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Molecular weight | Experimental: 150 MDa |
| Virus shell | Shell ID: 1 / Name: capsid / Diameter: 920 Å / T number (triangulation number): 25 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 / Details: Phosphate-buffered Saline (PBS) |
|---|---|
| Grid | Details: 200 mesh copper grid |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 77 K / Instrument: OTHER / Details: Vitrification instrument: Vitrobot / Method: Blotted twice for one second each. |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 2010F |
|---|---|
| Temperature | Min: 77 K |
| Date | Dec 17, 2004 |
| Image recording | Category: CCD / Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Sampling interval: 2.17 µm / Number real images: 100 / Average electron dose: 15 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 18.0 µm / Nominal defocus min: 0.5 µm / Nominal magnification: 50000 |
| Sample stage | Specimen holder: Side entry liquid nitrogen-cooled cryo specimen holder.This holder operates in the temperature range from 77K to ambient. Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| CTF correction | Details: Each micrograph |
|---|---|
| Final reconstruction | Applied symmetry - Point group: I (icosahedral) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 22.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: SAVR / Number images used: 800 |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















