[English] 日本語
 Yorodumi
Yorodumi- EMDB-1269: Structure of bacteriophage SPP1 tail reveals trigger for DNA ejection. -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1269 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of bacteriophage SPP1 tail reveals trigger for DNA ejection. | |||||||||
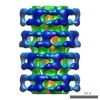
 Map data Map data | ccp4 map of the tip of the bacteriophage spp1 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Bacillus phage SPP1 (virus) Bacillus phage SPP1 (virus) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 20.0 Å | |||||||||
 Authors Authors | Plisson C / White HE / Auzat I / Sao-jose C / Lhuillier S / Tavares P / Orlova EV | |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2007 Journal: EMBO J / Year: 2007Title: Structure of bacteriophage SPP1 tail reveals trigger for DNA ejection. Authors: Celia Plisson / Helen E White / Isabelle Auzat / Amineh Zafarani / Carlos São-José / Sophie Lhuillier / Paulo Tavares / Elena V Orlova /  Abstract: The majority of known bacteriophages have long noncontractile tails (Siphoviridae) that serve as a pipeline for genome delivery into the host cytoplasm. The tail extremity distal from the phage head ...The majority of known bacteriophages have long noncontractile tails (Siphoviridae) that serve as a pipeline for genome delivery into the host cytoplasm. The tail extremity distal from the phage head is an adsorption device that recognises the bacterial receptor at the host cell surface. This interaction generates a signal transmitted to the head that leads to DNA release. We have determined structures of the bacteriophage SPP1 tail before and after DNA ejection. The results reveal extensive structural rearrangements in the internal wall of the tail tube. We propose that the adsorption device-receptor interaction triggers a conformational switch that is propagated as a domino-like cascade along the 1600 A-long helical tail structure to reach the head-to-tail connector. This leads to opening of the connector culminating in DNA exit from the head into the host cell through the tail tube. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1269.map.gz emd_1269.map.gz | 124.6 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1269-v30.xml emd-1269-v30.xml emd-1269.xml emd-1269.xml | 9.7 KB 9.7 KB | Display Display |  EMDB header EMDB header |
| Images |  1269.gif 1269.gif | 18.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1269 http://ftp.pdbj.org/pub/emdb/structures/EMD-1269 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1269 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1269 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1269.map.gz / Format: CCP4 / Size: 3.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1269.map.gz / Format: CCP4 / Size: 3.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | ccp4 map of the tip of the bacteriophage spp1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.3 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Tip of the Tail of the bacteriophage SPP1
| Entire | Name: Tip of the Tail of the bacteriophage SPP1 |
|---|---|
| Components |
|
-Supramolecule #1000: Tip of the Tail of the bacteriophage SPP1
| Supramolecule | Name: Tip of the Tail of the bacteriophage SPP1 / type: sample / ID: 1000 Details: The total number of proteins that compose the tail Tip is still under investigation. Number unique components: 2 |
|---|
-Macromolecule #1: interface tail-tip
| Macromolecule | Name: interface tail-tip / type: protein_or_peptide / ID: 1 / Name.synonym: gp19.1 / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  Bacillus phage SPP1 (virus) / synonym: Bacteriophage SPP1 Bacillus phage SPP1 (virus) / synonym: Bacteriophage SPP1 |
| Molecular weight | Experimental: 28.5 KDa |
-Macromolecule #2: tip fiber
| Macromolecule | Name: tip fiber / type: protein_or_peptide / ID: 2 / Name.synonym: gp21 / Recombinant expression: No |
|---|---|
| Source (natural) | Organism:  Bacillus phage SPP1 (virus) / synonym: Bacteriophage SPP1 Bacillus phage SPP1 (virus) / synonym: Bacteriophage SPP1 |
| Molecular weight | Experimental: 123 KDa |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 / Details: 100 mM Tris-Cl, 100 mM NaCl, 10 mM MgCl2 |
|---|---|
| Staining | Type: NEGATIVE Details: 3.5 microliter of sample was applied onto carbon-coated copper grids, which were glow-discharged for 30 seconds. The sample was left for 1-2 minutes on the grids before blotting and staining ...Details: 3.5 microliter of sample was applied onto carbon-coated copper grids, which were glow-discharged for 30 seconds. The sample was left for 1-2 minutes on the grids before blotting and staining with a solution of 2% uranyl acetate. |
| Grid | Details: 300 mesh copper grid |
| Vitrification | Cryogen name: NONE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Temperature | Average: 298 K |
| Alignment procedure | Legacy - Astigmatism: objective lens astigmatism was corrected at 150,000 times magnification |
| Details | Low Dose Imaging |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 7 µm / Number real images: 20 / Average electron dose: 10 e/Å2 / Od range: 2.5 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | Calibrated magnification: 43475 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 42000 |
| Sample stage | Specimen holder: Gatan single tilt negative stain holder / Specimen holder model: OTHER |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 20.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: IMAGIC Details: The 3D reconstruction has been caried out using a C3 symmetry and the exact-filter back projection method. Number images used: 364 |
|---|
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera



 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















