[English] 日本語
 Yorodumi
Yorodumi- EMDB-12224: Vps26 dimer region of the fungal membrane-assembled retromer:Grd1... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12224 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Vps26 dimer region of the fungal membrane-assembled retromer:Grd19 complex. | ||||||||||||||||||
 Map data Map data | Sharpened, locally filtered map of VPS26 dimer region of the fungal retromer:Grd19 complex assembled on the membrane | ||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | endosomes / coat proteins / membrane trafficking / cargo-sorting / ENDOCYTOSIS | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationvanadium ion transmembrane transporter activity / vanadium ion transport / paraferritin complex / Defective SLC11A2 causes hypochromic microcytic anemia, with iron overload 1 (AHMIO1) / lead ion transmembrane transporter activity / lead ion transport / nickel cation transmembrane transporter activity / transition metal ion transmembrane transporter activity / late endosome to Golgi transport / solute:proton symporter activity ...vanadium ion transmembrane transporter activity / vanadium ion transport / paraferritin complex / Defective SLC11A2 causes hypochromic microcytic anemia, with iron overload 1 (AHMIO1) / lead ion transmembrane transporter activity / lead ion transport / nickel cation transmembrane transporter activity / transition metal ion transmembrane transporter activity / late endosome to Golgi transport / solute:proton symporter activity / cadmium ion transmembrane transport / : / Metal ion SLC transporters / manganese ion transport / cadmium ion transmembrane transporter activity / detection of oxygen / nickel cation transport / retromer, cargo-selective complex / manganese ion transmembrane transporter activity / copper ion transmembrane transporter activity / cobalt ion transport / iron import into cell / intracellular manganese ion homeostasis / cobalt ion transmembrane transporter activity / retromer complex binding / ferrous iron transmembrane transporter activity / iron ion transmembrane transporter activity / zinc ion transmembrane transporter activity / retromer complex / iron ion transmembrane transport / copper ion transport / basal part of cell / phosphatidylinositol-3-phosphate binding / endocytic recycling / vacuole / retrograde transport, endosome to Golgi / response to iron ion / heme biosynthetic process / dendrite morphogenesis / cadmium ion binding / erythrocyte development / trans-Golgi network / iron ion transport / brush border membrane / Iron uptake and transport / intracellular protein transport / multicellular organismal-level iron ion homeostasis / recycling endosome / recycling endosome membrane / apical part of cell / late endosome / late endosome membrane / protein transport / extracellular vesicle / cellular response to oxidative stress / cytoplasmic vesicle / early endosome membrane / intracellular iron ion homeostasis / learning or memory / response to hypoxia / early endosome / mitochondrial outer membrane / lysosome / endosome membrane / apical plasma membrane / Golgi membrane / lysosomal membrane / perinuclear region of cytoplasm / cell surface / Golgi apparatus / mitochondrion / membrane / nucleus / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||||||||||||||
| Biological species |  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) / Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) /  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) | ||||||||||||||||||
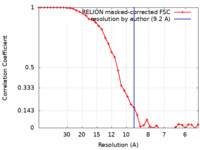
| Method | subtomogram averaging / cryo EM / Resolution: 9.2 Å | ||||||||||||||||||
 Authors Authors | Leneva N / Kovtun O | ||||||||||||||||||
| Funding support |  United Kingdom, 5 items United Kingdom, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Sci Adv / Year: 2021 Journal: Sci Adv / Year: 2021Title: Architecture and mechanism of metazoan retromer:SNX3 tubular coat assembly. Authors: Natalya Leneva / Oleksiy Kovtun / Dustin R Morado / John A G Briggs / David J Owen /  Abstract: Retromer is a master regulator of cargo retrieval from endosomes, which is critical for many cellular processes including signaling, immunity, neuroprotection, and virus infection. The retromer core ...Retromer is a master regulator of cargo retrieval from endosomes, which is critical for many cellular processes including signaling, immunity, neuroprotection, and virus infection. The retromer core (VPS26/VPS29/VPS35) is present on cargo-transporting, tubular carriers along with a range of sorting nexins. Here, we elucidate the structural basis of membrane tubulation and coupled cargo recognition by metazoan and fungal retromer coats assembled with the non-Bin1/Amphiphysin/Rvs (BAR) sorting nexin SNX3 using cryo-electron tomography. The retromer core retains its arched, scaffolding structure but changes its mode of membrane recruitment when assembled with different SNX adaptors, allowing cargo recognition at subunit interfaces. Thus, membrane bending and cargo incorporation can be modulated to allow retromer to traffic cargoes along different cellular transport routes. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12224.map.gz emd_12224.map.gz | 2.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12224-v30.xml emd-12224-v30.xml emd-12224.xml emd-12224.xml | 18.8 KB 18.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12224_fsc.xml emd_12224_fsc.xml | 3.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_12224.png emd_12224.png | 98.3 KB | ||
| Masks |  emd_12224_msk_1.map emd_12224_msk_1.map | 3.8 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-12224.cif.gz emd-12224.cif.gz | 6.1 KB | ||
| Others |  emd_12224_half_map_1.map.gz emd_12224_half_map_1.map.gz emd_12224_half_map_2.map.gz emd_12224_half_map_2.map.gz | 3 MB 3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12224 http://ftp.pdbj.org/pub/emdb/structures/EMD-12224 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12224 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12224 | HTTPS FTP |
-Related structure data
| Related structure data |  7blqMC  7blnC  7bloC  7blpC  7blrC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10631 (Title: Cryo-electron tomography of the fungal membrane-assembled retromer:Grd19 coat containing Kex2 cargo motif EMPIAR-10631 (Title: Cryo-electron tomography of the fungal membrane-assembled retromer:Grd19 coat containing Kex2 cargo motifData size: 347.7 Data #1: Raw image frames for the fungal retromer:Grd19 coat assembled on the Kex2 cargo-containing membranes [micrographs - multiframe] Data #2: Corrected, aligned and order-sorted tilt series for the membrane-reconstituted fungal retromer:Grd19 complex in the presence of cargo-signal containing C-portion of the Kex2 cargo. [tilt series] Data #3: Corrected, aligned, dose-filtered and order-sorted tilt series for the membrane-reconstituted fungal retromer:Grd19 complex in the presence of cargo-signal containing C-portion of the Kex2 cargo. [tilt series]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12224.map.gz / Format: CCP4 / Size: 3.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12224.map.gz / Format: CCP4 / Size: 3.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpened, locally filtered map of VPS26 dimer region of the fungal retromer:Grd19 complex assembled on the membrane | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.758 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12224_msk_1.map emd_12224_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half-map2
| File | emd_12224_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half-map1
| File | emd_12224_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half-map1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Vps26 dimer region of the fungal membrane-assembled retromer:Grd1...
| Entire | Name: Vps26 dimer region of the fungal membrane-assembled retromer:Grd19 cargo-containing complex |
|---|---|
| Components |
|
-Supramolecule #1: Vps26 dimer region of the fungal membrane-assembled retromer:Grd1...
| Supramolecule | Name: Vps26 dimer region of the fungal membrane-assembled retromer:Grd19 cargo-containing complex type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 Details: fungal retromer:Grd19 complex assembled on liposomes containing Kex2 cargo peptide. |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) |
-Macromolecule #1: Vacuolar protein sorting-associated protein 35
| Macromolecule | Name: Vacuolar protein sorting-associated protein 35 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus)Strain: DSM 1495 / CBS 144.50 / IMI 039719 |
| Molecular weight | Theoretical: 34.014195 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: RLLEDALIAV RQQTAMMRKF LDTPGKLMDA LKCCSTLVSE LRTSSLSPKQ YYELYMAVFD ALRYLSAHLR ENHPVNHLAD LYELVQYAG NIIPRLYLMI TVGTAYMSID GAPVKELMKD MMDMSRGVQH PVRGLFLRYY LSGQARDYLP TGDSDGPEGN L QDSINFIL ...String: RLLEDALIAV RQQTAMMRKF LDTPGKLMDA LKCCSTLVSE LRTSSLSPKQ YYELYMAVFD ALRYLSAHLR ENHPVNHLAD LYELVQYAG NIIPRLYLMI TVGTAYMSID GAPVKELMKD MMDMSRGVQH PVRGLFLRYY LSGQARDYLP TGDSDGPEGN L QDSINFIL TNFVEMNKLW VRLQHQGHSR ERDLRTQERR ELQLLVGSNI VRLSQLVDLP TYRDSILGPL LEQIVQCRDI LA QEYLLEV ITQVFPDEYH LHTLDQFLGA VSRLNPHVNV KAIVIGMMNR LSDYAERE UniProtKB: Vacuolar protein sorting-associated protein 35 |
-Macromolecule #2: Sorting nexin-3
| Macromolecule | Name: Sorting nexin-3 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus)Strain: DSM 1495 / CBS 144.50 / IMI 039719 |
| Molecular weight | Theoretical: 13.52549 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: PPENFLEIEV RNPQTHGVGR HMYTDYEIVC RTNIPAFKLR QSSVRRRYSD FEYFRDILER ESARVTIPPL PGKVFTNRFS DEVIENRRA GLEKFLKIVV GHPLLQTGSK VLAAFVQ UniProtKB: Sorting nexin-3 |
-Macromolecule #3: Vacuolar protein sorting-associated protein 26-like protein
| Macromolecule | Name: Vacuolar protein sorting-associated protein 26-like protein type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (fungus)Strain: DSM 1495 / CBS 144.50 / IMI 039719 |
| Molecular weight | Theoretical: 34.308449 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: FSTPVDIDIV LADADKRAMV DVKLDKNRRE KVPLYMDGES VKGCVTVRPK DGKRLEHTGI KVQFIGTIEM FFDRGNHYEF LSLVQELAA PGELQHPQTF DFNFKNVEKQ YESYNGINVK LRYFVRVTVS RRMADVIREK DIWVYSYRIP PELNSSIKMD V GIEDCLHI ...String: FSTPVDIDIV LADADKRAMV DVKLDKNRRE KVPLYMDGES VKGCVTVRPK DGKRLEHTGI KVQFIGTIEM FFDRGNHYEF LSLVQELAA PGELQHPQTF DFNFKNVEKQ YESYNGINVK LRYFVRVTVS RRMADVIREK DIWVYSYRIP PELNSSIKMD V GIEDCLHI EFEYSKSKYH LKDVIVGRIY FLLVRLKIKH MELSIIRRET TGVAPNQYNE SETLVRFEIM DGSPSRGETI PI RLFLGGF DLTPTFRDVN KKFSTRYYLS LVLIDEDARR YFKQSEIILY RQPPE UniProtKB: Vacuolar protein sorting-associated protein 26-like protein |
-Macromolecule #4: The C-terminal portion of Kex2 cargo, fitted with Phi-X-(L/M) sor...
| Macromolecule | Name: The C-terminal portion of Kex2 cargo, fitted with Phi-X-(L/M) sorting motif of hDMT1-II cargo. type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) Chaetomium thermophilum var. thermophilum DSM 1495 (fungus) |
| Molecular weight | Theoretical: 1.221422 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: QPELYLLNTM |
-Macromolecule #5: 2-(BUTANOYLOXY)-1-{[(HYDROXY{[2,3,4,6-TETRAHYDROXY-5-(PHOSPHONOOX...
| Macromolecule | Name: 2-(BUTANOYLOXY)-1-{[(HYDROXY{[2,3,4,6-TETRAHYDROXY-5-(PHOSPHONOOXY)CYCLOHEXYL]OXY}PHOSPHORYL)OXY]METHYL}ETHYL BUTANOATE type: ligand / ID: 5 / Number of copies: 2 / Formula: PIB |
|---|---|
| Molecular weight | Theoretical: 554.374 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | 3D array |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 3.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)