[English] 日本語
 Yorodumi
Yorodumi- EMDB-11802: Structure of Human Potassium Chloride Transporter KCC1 in NaCl (S... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11802 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Human Potassium Chloride Transporter KCC1 in NaCl (Subclass 1) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CCC / transporter / human membrane protein / homodimer / nucleotide-binding / KCC1 / KCC / potassium-chloride coupled transporter / Structural Genomics / Structural Genomics Consortium / SGC / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationpotassium:chloride symporter activity / Cation-coupled Chloride cotransporters / chloride ion homeostasis / potassium ion homeostasis / cell volume homeostasis / potassium ion import across plasma membrane / monoatomic ion transport / potassium ion transmembrane transport / chloride transmembrane transport / protein serine/threonine kinase binding ...potassium:chloride symporter activity / Cation-coupled Chloride cotransporters / chloride ion homeostasis / potassium ion homeostasis / cell volume homeostasis / potassium ion import across plasma membrane / monoatomic ion transport / potassium ion transmembrane transport / chloride transmembrane transport / protein serine/threonine kinase binding / chemical synaptic transmission / lysosomal membrane / synapse / ATP binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
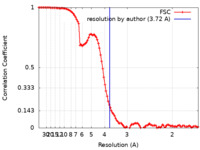
| Method | single particle reconstruction / cryo EM / Resolution: 3.72 Å | |||||||||
 Authors Authors | Ebenhoch R / Chi G | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2021 Journal: EMBO J / Year: 2021Title: Phospho-regulation, nucleotide binding and ion access control in potassium-chloride cotransporters. Authors: Gamma Chi / Rebecca Ebenhoch / Henry Man / Haiping Tang / Laurence E Tremblay / Gabriella Reggiano / Xingyu Qiu / Tina Bohstedt / Idlir Liko / Fernando G Almeida / Alexandre P Garneau / Dong ...Authors: Gamma Chi / Rebecca Ebenhoch / Henry Man / Haiping Tang / Laurence E Tremblay / Gabriella Reggiano / Xingyu Qiu / Tina Bohstedt / Idlir Liko / Fernando G Almeida / Alexandre P Garneau / Dong Wang / Gavin McKinley / Christophe P Moreau / Kiran D Bountra / Patrizia Abrusci / Shubhashish M M Mukhopadhyay / Alejandra Fernandez-Cid / Samira Slimani / Julie L Lavoie / Nicola A Burgess-Brown / Ben Tehan / Frank DiMaio / Ali Jazayeri / Paul Isenring / Carol V Robinson / Katharina L Dürr /    Abstract: Potassium-coupled chloride transporters (KCCs) play crucial roles in regulating cell volume and intracellular chloride concentration. They are characteristically inhibited under isotonic conditions ...Potassium-coupled chloride transporters (KCCs) play crucial roles in regulating cell volume and intracellular chloride concentration. They are characteristically inhibited under isotonic conditions via phospho-regulatory sites located within the cytoplasmic termini. Decreased inhibitory phosphorylation in response to hypotonic cell swelling stimulates transport activity, and dysfunction of this regulatory process has been associated with various human diseases. Here, we present cryo-EM structures of human KCC3b and KCC1, revealing structural determinants for phospho-regulation in both N- and C-termini. We show that phospho-mimetic KCC3b is arrested in an inward-facing state in which intracellular ion access is blocked by extensive contacts with the N-terminus. In another mutant with increased isotonic transport activity, KCC1Δ19, this interdomain interaction is absent, likely due to a unique phospho-regulatory site in the KCC1 N-terminus. Furthermore, we map additional phosphorylation sites as well as a previously unknown ATP/ADP-binding pocket in the large C-terminal domain and show enhanced thermal stabilization of other CCCs by adenine nucleotides. These findings provide fundamentally new insights into the complex regulation of KCCs and may unlock innovative strategies for drug development. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11802.map.gz emd_11802.map.gz | 97.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11802-v30.xml emd-11802-v30.xml emd-11802.xml emd-11802.xml | 17.3 KB 17.3 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11802_fsc.xml emd_11802_fsc.xml | 11.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_11802.png emd_11802.png | 163.6 KB | ||
| Masks |  emd_11802_msk_1.map emd_11802_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-11802.cif.gz emd-11802.cif.gz | 6.7 KB | ||
| Others |  emd_11802_half_map_1.map.gz emd_11802_half_map_1.map.gz emd_11802_half_map_2.map.gz emd_11802_half_map_2.map.gz | 95.7 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11802 http://ftp.pdbj.org/pub/emdb/structures/EMD-11802 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11802 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11802 | HTTPS FTP |
-Related structure data
| Related structure data |  7aiqMC  6y5vC  7ainC  7aioC  7aipC  7airC  7ngbC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11802.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11802.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11802_msk_1.map emd_11802_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_11802_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_11802_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Homodimeric complex of human potassium chloride transporter KCC1 ...
| Entire | Name: Homodimeric complex of human potassium chloride transporter KCC1 in complex with ATP and magnesium |
|---|---|
| Components |
|
-Supramolecule #1: Homodimeric complex of human potassium chloride transporter KCC1 ...
| Supramolecule | Name: Homodimeric complex of human potassium chloride transporter KCC1 in complex with ATP and magnesium type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 240 KDa |
-Macromolecule #1: Solute carrier family 12 member 4
| Macromolecule | Name: Solute carrier family 12 member 4 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 119.610281 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MLEGLSWVDY GERAELDDSD GHGNHRESSP FLSPLEASRG IDYYDRNLAL FEEELDIRPK VSSLLGKLVS YTNLTQGAKE HEEAESGEG TRRRAAEAPS MGTLMGVYLP CLQNIFGVIL FLRLTWMVGT AGVLQALLIV LICCCCTLLT AISMSAIATN G VVPAGGSY ...String: MLEGLSWVDY GERAELDDSD GHGNHRESSP FLSPLEASRG IDYYDRNLAL FEEELDIRPK VSSLLGKLVS YTNLTQGAKE HEEAESGEG TRRRAAEAPS MGTLMGVYLP CLQNIFGVIL FLRLTWMVGT AGVLQALLIV LICCCCTLLT AISMSAIATN G VVPAGGSY FMISRSLGPE FGGAVGLCFY LGTTFAAAMY ILGAIEILLT YIAPPAAIFY PSGAHDTSNA TLNNMRVYGT IF LTFMTLV VFVGVKYVNK FASLFLACVI ISILSIYAGG IKSIFDPPVF PVCMLGNRTL SRDQFDICAK TAVVDNETVA TQL WSFFCH SPNLTTDSCD PYFMLNNVTE IPGIPGAAAG VLQENLWSAY LEKGDIVEKH GLPSADAPSL KESLPLYVVA DIAT SFTVL VGIFFPSVTG IMAGSNRSGD LRDAQKSIPV GTILAIITTS LVYFSSVVLF GACIEGVVLR DKYGDGVSRN LVVGT LAWP SPWVIVIGSF FSTCGAGLQS LTGAPRLLQA IAKDNIIPFL RVFGHGKVNG EPTWALLLTA LIAELGILIA SLDMVA PIL SMFFLMCYLF VNLACAVQTL LRTPNWRPRF KYYHWALSFL GMSLCLALMF VSSWYYALVA MLIAGMIYKY IEYQGAE KE WGDGIRGLSL SAARYALLRL EEGPPHTKNW RPQLLVLLKL DEDLHVKYPR LLTFASQLKA GKGLTIVGSV IQGSFLES Y GEAQAAEQTI KNMMEIEKVK GFCQVVVASK VREGLAHLIQ SCGLGGMRHN SVVLGWPYGW RQSEDPRAWK TFIDTVRCT TAAHLALLVP KNIAFYPSNH ERYLEGHIDV WWIVHDGGML MLLPFLLRQH KVWRKCRMRI FTVAQMDDNS IQMKKDLAVF LYHLRLEAE VEVVEMHNSD ISAYTYERTL MMEQRSQMLR QMRLTKTERE REAQLVKDRH SALRLESLYS DEEDESAVGA D KIQMTWTR DKYMTETWDP SHAPDNFREL VHIKPDQSNV RRMHTAVKLN EVIVTRSHDA RLVLLNMPGP PRNSEGDENY ME FLEVLTE GLERVLLVRG GGREVITIYS AENLYFQ UniProtKB: Solute carrier family 12 member 4 |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #4: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 4 / Number of copies: 2 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Model from reference map/model fitted with Chimera, refined with Coot and then with Phenix. |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
| Output model |  PDB-7aiq: |
 Movie
Movie Controller
Controller


















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)