+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11085 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
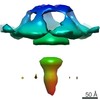
| Title | Radial spoke head structure with 2-fold symmetization | |||||||||
 Map data Map data | This is the map of the radial spoke from cilia with 2-fold symmetry applied. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 15.0 Å | |||||||||
 Authors Authors | Poghosyan E / Ishikawa T | |||||||||
| Funding support |  Switzerland, 1 items Switzerland, 1 items
| |||||||||
 Citation Citation |  Journal: J Cell Sci / Year: 2020 Journal: J Cell Sci / Year: 2020Title: The structure and symmetry of the radial spoke protein complex in flagella. Authors: Emiliya Poghosyan / Ioan Iacovache / Lenka Faltova / Alexander Leitner / Pinfen Yang / Dennis R Diener / Ruedi Aebersold / Benoit Zuber / Takashi Ishikawa /   Abstract: The radial spoke is a key element in a transducer apparatus controlling the motility of eukaryotic cilia. The transduction biomechanics is a long-standing question in cilia biology. The radial spoke ...The radial spoke is a key element in a transducer apparatus controlling the motility of eukaryotic cilia. The transduction biomechanics is a long-standing question in cilia biology. The radial spoke has three regions - a spoke head, a bifurcated neck and a stalk. Although the neck and the stalk are asymmetric, twofold symmetry of the head has remained controversial. In this work we used single particle cryo-electron microscopy (cryo-EM) analysis to generate a 3D structure of the whole radial spoke at unprecedented resolution. We show the head region at 15 Å (1.5 nm) resolution and confirm twofold symmetry. Using distance constraints generated by cross-linking mass spectrometry, we locate two components, RSP2 and RSP4, at the head and neck regions. Our biophysical analysis of isolated RSP4, RSP9, and RSP10 affirmed their oligomeric state. Our results enable us to redefine the boundaries of the regions and propose a model of organization of the radial spoke component proteins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11085.map.gz emd_11085.map.gz | 13.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11085-v30.xml emd-11085-v30.xml emd-11085.xml emd-11085.xml | 10.9 KB 10.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11085.png emd_11085.png | 85.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11085 http://ftp.pdbj.org/pub/emdb/structures/EMD-11085 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11085 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11085 | HTTPS FTP |
-Validation report
| Summary document |  emd_11085_validation.pdf.gz emd_11085_validation.pdf.gz | 199.8 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_11085_full_validation.pdf.gz emd_11085_full_validation.pdf.gz | 198.9 KB | Display | |
| Data in XML |  emd_11085_validation.xml.gz emd_11085_validation.xml.gz | 6.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11085 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11085 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11085 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-11085 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11085.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11085.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | This is the map of the radial spoke from cilia with 2-fold symmetry applied. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 2.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Radial spoke of cilia
| Entire | Name: Radial spoke of cilia |
|---|---|
| Components |
|
-Supramolecule #1: Radial spoke of cilia
| Supramolecule | Name: Radial spoke of cilia / type: complex / ID: 1 / Parent: 0 Details: Radial spoke extracted from Chlamydomonas flagella/cilia |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 1.2 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 10 mM HEPES pH 7.5, 5 mM MgSO4, 1 mM BME, 0.5 mM EDTA, 25 mM KCl |
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 5.0 nm / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 20 K / Instrument: FEI VITROBOT MARK III |
| Details | Solution of protein complexes isolated from natural source |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 2 / Number real images: 736 / Average exposure time: 7.0 sec. / Average electron dose: 20.0 e/Å2 / Details: 7 frames/sec |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm |
| Sample stage | Specimen holder model: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)





















