[English] 日本語
 Yorodumi
Yorodumi- EMDB-10717: head domain of Tsr1-FPZ pre-40S particle: structural class 2 (H2) -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10717 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | head domain of Tsr1-FPZ pre-40S particle: structural class 2 (H2) | |||||||||
 Map data Map data | H2-Tsr1 pre-40S Head of yeast Tsr1-FPZ pre-40S particle Structural class 2 | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
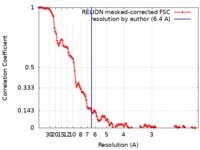
| Method | single particle reconstruction / cryo EM / Resolution: 6.4 Å | |||||||||
 Authors Authors | Shayan R / Plassart L / Plisson-Chastang C | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Molecules / Year: 2020 Journal: Molecules / Year: 2020Title: Good Vibrations: Structural Remodeling of Maturing Yeast Pre-40S Ribosomal Particles Followed by Cryo-Electron Microscopy. Authors: Ramtin Shayan / Dana Rinaldi / Natacha Larburu / Laura Plassart / Stéphanie Balor / David Bouyssié / Simon Lebaron / Julien Marcoux / Pierre-Emmanuel Gleizes / Célia Plisson-Chastang /  Abstract: Assembly of eukaryotic ribosomal subunits is a very complex and sequential process that starts in the nucleolus and finishes in the cytoplasm with the formation of functional ribosomes. Over the past ...Assembly of eukaryotic ribosomal subunits is a very complex and sequential process that starts in the nucleolus and finishes in the cytoplasm with the formation of functional ribosomes. Over the past few years, characterization of the many molecular events underlying eukaryotic ribosome biogenesis has been drastically improved by the "resolution revolution" of cryo-electron microscopy (cryo-EM). However, if very early maturation events have been well characterized for both yeast ribosomal subunits, little is known regarding the final maturation steps occurring to the small (40S) ribosomal subunit. To try to bridge this gap, we have used proteomics together with cryo-EM and single particle analysis to characterize yeast pre-40S particles containing the ribosome biogenesis factor Tsr1. Our analyses lead us to refine the timing of the early pre-40S particle maturation steps. Furthermore, we suggest that after an early and structurally stable stage, the beak and platform domains of pre-40S particles enter a "vibrating" or "wriggling" stage, that might be involved in the final maturation of 18S rRNA as well as the fitting of late ribosomal proteins into their mature position. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10717.map.gz emd_10717.map.gz | 13.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10717-v30.xml emd-10717-v30.xml emd-10717.xml emd-10717.xml | 20 KB 20 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10717_fsc.xml emd_10717_fsc.xml | 12.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_10717.png emd_10717.png | 123.5 KB | ||
| Others |  emd_10717_half_map_1.map.gz emd_10717_half_map_1.map.gz emd_10717_half_map_2.map.gz emd_10717_half_map_2.map.gz | 140.7 MB 140.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10717 http://ftp.pdbj.org/pub/emdb/structures/EMD-10717 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10717 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10717 | HTTPS FTP |
-Validation report
| Summary document |  emd_10717_validation.pdf.gz emd_10717_validation.pdf.gz | 321.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10717_full_validation.pdf.gz emd_10717_full_validation.pdf.gz | 320.8 KB | Display | |
| Data in XML |  emd_10717_validation.xml.gz emd_10717_validation.xml.gz | 19.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10717 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10717 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10717 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10717 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10717.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10717.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | H2-Tsr1 pre-40S Head of yeast Tsr1-FPZ pre-40S particle Structural class 2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.067 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: H2-Tsr1 pre-40S Head of yeast Tsr1-FPZ pre-40S particle...
| File | emd_10717_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | H2-Tsr1 pre-40S Head of yeast Tsr1-FPZ pre-40S particle Structural class 2 Unfiltered half-map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: H2-Tsr1 pre-40S Head of yeast Tsr1-FPZ pre-40S particle...
| File | emd_10717_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | H2-Tsr1 pre-40S Head of yeast Tsr1-FPZ pre-40S particle Structural class 2 Unfiltered half-map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : early cytoplasmic yeast pre-40S particle (purified using Tsr1 as bait)
| Entire | Name: early cytoplasmic yeast pre-40S particle (purified using Tsr1 as bait) |
|---|---|
| Components |
|
-Supramolecule #1: early cytoplasmic yeast pre-40S particle (purified using Tsr1 as bait)
| Supramolecule | Name: early cytoplasmic yeast pre-40S particle (purified using Tsr1 as bait) type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#34 |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 1.8 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: CONTINUOUS / Support film - Film thickness: 2.0 nm / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 292 K / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 29.4 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Refinement | Protocol: OTHER |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)