[English] 日本語
 Yorodumi
Yorodumi- EMDB-10707: Anabaena sp. PCC 7120 Septal Junction Average from Manually Mille... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10707 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Anabaena sp. PCC 7120 Septal Junction Average from Manually Milled Lamellae | |||||||||
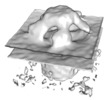
 Map data Map data | Septal junction average generated from 343 particles and 5-fold symmetrizing. Lamellae were generated in using a manual FIB milling approach. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Nostoc sp. PCC 7120 (bacteria) Nostoc sp. PCC 7120 (bacteria) | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 30.0 Å | |||||||||
 Authors Authors | Zachs T / Pilhofer M | |||||||||
| Funding support |  Switzerland, European Union, 2 items Switzerland, European Union, 2 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: Fully automated, sequential focused ion beam milling for cryo-electron tomography. Authors: Tobias Zachs / Andreas Schertel / João Medeiros / Gregor L Weiss / Jannik Hugener / Joao Matos / Martin Pilhofer /   Abstract: Cryo-electron tomography (cryoET) has become a powerful technique at the interface of structural biology and cell biology, due to its unique ability for imaging cells in their native state and ...Cryo-electron tomography (cryoET) has become a powerful technique at the interface of structural biology and cell biology, due to its unique ability for imaging cells in their native state and determining structures of macromolecular complexes in their cellular context. A limitation of cryoET is its restriction to relatively thin samples. Sample thinning by cryo-focused ion beam (cryoFIB) milling has significantly expanded the range of samples that can be analyzed by cryoET. Unfortunately, cryoFIB milling is low-throughput, time-consuming and manual. Here, we report a method for fully automated sequential cryoFIB preparation of high-quality lamellae, including rough milling and polishing. We reproducibly applied this method to eukaryotic and bacterial model organisms, and show that the resulting lamellae are suitable for cryoET imaging and subtomogram averaging. Since our method reduces the time required for lamella preparation and minimizes the need for user input, we envision the technique will render previously inaccessible projects feasible. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10707.map.gz emd_10707.map.gz | 273 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10707-v30.xml emd-10707-v30.xml emd-10707.xml emd-10707.xml | 7.7 KB 7.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10707.png emd_10707.png | 61.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10707 http://ftp.pdbj.org/pub/emdb/structures/EMD-10707 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10707 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10707 | HTTPS FTP |
-Validation report
| Summary document |  emd_10707_validation.pdf.gz emd_10707_validation.pdf.gz | 225 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_10707_full_validation.pdf.gz emd_10707_full_validation.pdf.gz | 224.1 KB | Display | |
| Data in XML |  emd_10707_validation.xml.gz emd_10707_validation.xml.gz | 5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10707 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10707 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10707 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-10707 | HTTPS FTP |
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10376 (Title: Yeast Tilt Series Collected on Lamella Generated by Fully Automated FIB Milling EMPIAR-10376 (Title: Yeast Tilt Series Collected on Lamella Generated by Fully Automated FIB MillingData size: 2.6 Data #1: Yeast Tilt Series Collected on Lamella Generated by Fully Automated FIB Milling [tilt series]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10707.map.gz / Format: CCP4 / Size: 333 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10707.map.gz / Format: CCP4 / Size: 333 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Septal junction average generated from 343 particles and 5-fold symmetrizing. Lamellae were generated in using a manual FIB milling approach. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 6.757 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
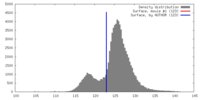
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Septal Junction
| Entire | Name: Septal Junction |
|---|---|
| Components |
|
-Supramolecule #1: Septal Junction
| Supramolecule | Name: Septal Junction / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Nostoc sp. PCC 7120 (bacteria) Nostoc sp. PCC 7120 (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 2.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C5 (5 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 30.0 Å / Resolution method: FSC 0.5 CUT-OFF / Number subtomograms used: 343 |
|---|---|
| Extraction | Number tomograms: 9 / Number images used: 343 |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)