[English] 日本語
 Yorodumi
Yorodumi- EMDB-1041: Nucleotide-induced conformations in the neck region of dimeric ki... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1041 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Nucleotide-induced conformations in the neck region of dimeric kinesin. | |||||||||
 Map data Map data | rat kinesin monomer rk354 complexed to microtubules in the absence of nucleotide | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | helical reconstruction / cryo EM / negative staining / Resolution: 25.0 Å | |||||||||
 Authors Authors | Skiniotis G | |||||||||
 Citation Citation |  Journal: EMBO J / Year: 2003 Journal: EMBO J / Year: 2003Title: Nucleotide-induced conformations in the neck region of dimeric kinesin. Authors: Georgios Skiniotis / Thomas Surrey / Stephan Altmann / Heinz Gross / Young-Hwa Song / Eckhard Mandelkow / Andreas Hoenger /  Abstract: The neck region of kinesin constitutes a key component in the enzyme's walking mechanism. Here we applied cryoelectron microscopy and image reconstruction to investigate the location of the kinesin ...The neck region of kinesin constitutes a key component in the enzyme's walking mechanism. Here we applied cryoelectron microscopy and image reconstruction to investigate the location of the kinesin neck in dimeric and monomeric constructs complexed to microtubules. To this end we enhanced the visibility of this region by engineering an SH3 domain into the transition between neck linker and neck coiled coil. The resulting chimeric kinesin constructs remained functional as verified by physiology assays. In the presence of AMP-PNP the SH3 domains allowed us to identify the position of the neck in a well defined conformation and revealed its high flexibility in the absence of nucleotide. We show here the double-headed binding of dimeric kinesin along the same protofilament, which is characterized by the opposite directionality of neck linkers. In this configuration the neck coiled coil appears fully zipped. The position of the neck region in dimeric constructs is not affected by the presence of the tubulin C-termini as confirmed by subtilisin treatment of microtubules prior to motor decoration. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1041.map.gz emd_1041.map.gz | 2.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1041-v30.xml emd-1041-v30.xml emd-1041.xml emd-1041.xml | 10.3 KB 10.3 KB | Display Display |  EMDB header EMDB header |
| Images |  1041.gif 1041.gif | 66 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1041 http://ftp.pdbj.org/pub/emdb/structures/EMD-1041 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1041 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1041 | HTTPS FTP |
-Validation report
| Summary document |  emd_1041_validation.pdf.gz emd_1041_validation.pdf.gz | 249.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1041_full_validation.pdf.gz emd_1041_full_validation.pdf.gz | 248.4 KB | Display | |
| Data in XML |  emd_1041_validation.xml.gz emd_1041_validation.xml.gz | 5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1041 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1041 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1041 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1041 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1041.map.gz / Format: CCP4 / Size: 3.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1041.map.gz / Format: CCP4 / Size: 3.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | rat kinesin monomer rk354 complexed to microtubules in the absence of nucleotide | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 5.526 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
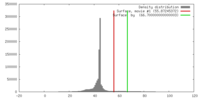
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : rat kinesin monomer ric354 complexed to microtubule in the absenc...
| Entire | Name: rat kinesin monomer ric354 complexed to microtubule in the absence of nucleotide |
|---|---|
| Components |
|
-Supramolecule #1000: rat kinesin monomer ric354 complexed to microtubule in the absenc...
| Supramolecule | Name: rat kinesin monomer ric354 complexed to microtubule in the absence of nucleotide type: sample / ID: 1000 / Oligomeric state: monomer / Number unique components: 2 |
|---|
-Macromolecule #1: rat kinesin
| Macromolecule | Name: rat kinesin / type: protein_or_peptide / ID: 1 / Name.synonym: molecular motor / Number of copies: 1 / Oligomeric state: dimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 390 KDa |
| Recombinant expression | Organism:  |
-Macromolecule #2: tubulin
| Macromolecule | Name: tubulin / type: protein_or_peptide / ID: 2 / Name.synonym: microtubule / Number of copies: 1 / Oligomeric state: hetero-dimer / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 110 KDa |
-Experimental details
-Structure determination
| Method | negative staining, cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 6.8 Details: PIPES 80 mM, MgCl2 1 mM, GTP 1 mM, Taxol 20 uM, DMSO 7.5% |
| Staining | Type: NEGATIVE / Details: ice-embedded |
| Grid | Details: holey grids |
| Vitrification | Cryogen name: ETHANE / Chamber temperature: 93 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI/PHILIPS CM200FEG/ST |
|---|---|
| Alignment procedure | Legacy - Astigmatism: was corrected at 180,000 times mag. |
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: ZEISS SCAI / Digitization - Sampling interval: 21 µm / Number real images: 10 / Average electron dose: 5 e/Å2 / Bits/pixel: 8 |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 2.0 µm / Nominal magnification: 38000 |
| Sample stage | Specimen holder: side entry / Specimen holder model: GATAN LIQUID NITROGEN |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Helical parameters - Axial symmetry: C1 (asymmetric) Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 25.0 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: PHOELIX, SUPRIM Details: Final map from 20 averaged datasets = 10 helical tubes |
|---|
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)