[English] 日本語
 Yorodumi
Yorodumi- EMDB-10409: In situ cryo-electron tomogram from Chlamydomonas reinhardtii of ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10409 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | In situ cryo-electron tomogram from Chlamydomonas reinhardtii of a proteasome cluster at the endoplasmic reticulum | |||||||||
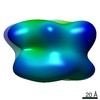
 Map data Map data | In situ cryo-electron tomogram from Chlamydomonas reinhardtii, showing a cluster of proteasomes at the ER membrane. | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
| Method | electron tomography / cryo EM | |||||||||
 Authors Authors | Albert S / Schaffer M / Baumeister W / Engel BD | |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2020 Journal: Proc Natl Acad Sci U S A / Year: 2020Title: Direct visualization of degradation microcompartments at the ER membrane. Authors: Sahradha Albert / Wojciech Wietrzynski / Chia-Wei Lee / Miroslava Schaffer / Florian Beck / Jan M Schuller / Patrice A Salomé / Jürgen M Plitzko / Wolfgang Baumeister / Benjamin D Engel /   Abstract: To promote the biochemical reactions of life, cells can compartmentalize molecular interaction partners together within separated non-membrane-bound regions. It is unknown whether this strategy is ...To promote the biochemical reactions of life, cells can compartmentalize molecular interaction partners together within separated non-membrane-bound regions. It is unknown whether this strategy is used to facilitate protein degradation at specific locations within the cell. Leveraging in situ cryo-electron tomography to image the native molecular landscape of the unicellular alga , we discovered that the cytosolic protein degradation machinery is concentrated within ∼200-nm foci that contact specialized patches of endoplasmic reticulum (ER) membrane away from the ER-Golgi interface. These non-membrane-bound microcompartments exclude ribosomes and consist of a core of densely clustered 26S proteasomes surrounded by a loose cloud of Cdc48. Active proteasomes in the microcompartments directly engage with putative substrate at the ER membrane, a function canonically assigned to Cdc48. Live-cell fluorescence microscopy revealed that the proteasome clusters are dynamic, with frequent assembly and fusion events. We propose that the microcompartments perform ER-associated degradation, colocalizing the degradation machinery at specific ER hot spots to enable efficient protein quality control. #1:  Journal: To Be Published Journal: To Be PublishedTitle: Direct visualization of degradation microcompartments at the ER membrane Authors: Albert S / Wietrzynski W / Lee CW / Schaffer M / Beck F / Schuller JM / Salome PA / Plitzko JM / Baumeister W / Engel BD | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10409.map.gz emd_10409.map.gz | 337 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10409-v30.xml emd-10409-v30.xml emd-10409.xml emd-10409.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_10409.png emd_10409.png | 185.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10409 http://ftp.pdbj.org/pub/emdb/structures/EMD-10409 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10409 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10409 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_10409.map.gz / Format: CCP4 / Size: 762.2 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) Download / File: emd_10409.map.gz / Format: CCP4 / Size: 762.2 MB / Type: IMAGE STORED AS SIGNED INTEGER (2 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | In situ cryo-electron tomogram from Chlamydomonas reinhardtii, showing a cluster of proteasomes at the ER membrane. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 13.68 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Whole Chlamydomonas cells
| Entire | Name: Whole Chlamydomonas cells |
|---|---|
| Components |
|
-Supramolecule #1: Whole Chlamydomonas cells
| Supramolecule | Name: Whole Chlamydomonas cells / type: cell / ID: 1 / Parent: 0 Details: Grown suspended in TAP media, with normal atmosphere aeration and constant light |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | electron tomography |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7 / Details: TAP media |
|---|---|
| Grid | Model: Quantifoil R2/1 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE-PROPANE / Instrument: FEI VITROBOT MARK IV / Details: blotted from back for 10 seconds before plunging. |
| Sectioning | Focused ion beam - Instrument: OTHER / Focused ion beam - Ion: OTHER / Focused ion beam - Voltage: 30 kV / Focused ion beam - Current: 0.03 nA / Focused ion beam - Duration: 2000 sec. / Focused ion beam - Temperature: 95 K / Focused ion beam - Initial thickness: 1000 nm / Focused ion beam - Final thickness: 150 nm Focused ion beam - Details: See https://bio-protocol.org/e1575 for detailed procedure.. The value given for _emd_sectioning_focused_ion_beam.instrument is FEI Quanta FIB. This is not in a list of ...Focused ion beam - Details: See https://bio-protocol.org/e1575 for detailed procedure.. The value given for _emd_sectioning_focused_ion_beam.instrument is FEI Quanta FIB. This is not in a list of allowed values set(['DB235', 'OTHER']) so OTHER is written into the XML file. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Details | Bidirectional tilt scheme, separated at 0 degrees |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average exposure time: 1.5 sec. / Average electron dose: 100.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 5.6 µm / Calibrated defocus min: 4.6 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 5.0 µm / Nominal defocus min: 5.0 µm / Nominal magnification: 42000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Algorithm: BACK PROJECTION / Software - Name:  IMOD / Number images used: 60 IMOD / Number images used: 60 |
|---|---|
| CTF correction | Software - Name:  IMOD IMOD |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)

















