+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10371 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of MexB in nanodisc | |||||||||
 Map data Map data | CryoEM structure of RND transport MexB trimer from Pseudomonas aeruginosa stabilized in nanodisc | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationxenobiotic detoxification by transmembrane export across the cell outer membrane / efflux pump complex / efflux transmembrane transporter activity / xenobiotic transmembrane transporter activity / transmembrane transport / response to toxic substance / response to antibiotic / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||
 Authors Authors | Glavier M / Puvanendran D / Salvador D / Decossas M / Phan G / Garnier C / Frezza E / Cece Q / Schoehn G / Picard M ...Glavier M / Puvanendran D / Salvador D / Decossas M / Phan G / Garnier C / Frezza E / Cece Q / Schoehn G / Picard M / Taveau J-C / Daury L / Broutin I / Lambert O | |||||||||
| Funding support |  France, 1 items France, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2020 Journal: Nat Commun / Year: 2020Title: Antibiotic export by MexB multidrug efflux transporter is allosterically controlled by a MexA-OprM chaperone-like complex. Authors: Marie Glavier / Dhenesh Puvanendran / Dimitri Salvador / Marion Decossas / Gilles Phan / Cyril Garnier / Elisa Frezza / Quentin Cece / Guy Schoehn / Martin Picard / Jean-Christophe Taveau / ...Authors: Marie Glavier / Dhenesh Puvanendran / Dimitri Salvador / Marion Decossas / Gilles Phan / Cyril Garnier / Elisa Frezza / Quentin Cece / Guy Schoehn / Martin Picard / Jean-Christophe Taveau / Laetitia Daury / Isabelle Broutin / Olivier Lambert /  Abstract: The tripartite multidrug efflux system MexAB-OprM is a major actor in Pseudomonas aeruginosa antibiotic resistance by exporting a large variety of antimicrobial compounds. Crystal structures of MexB ...The tripartite multidrug efflux system MexAB-OprM is a major actor in Pseudomonas aeruginosa antibiotic resistance by exporting a large variety of antimicrobial compounds. Crystal structures of MexB and of its Escherichia coli homolog AcrB had revealed asymmetric trimers depicting a directional drug pathway by a conformational interconversion (from Loose and Tight binding pockets to Open gate (LTO) for drug exit). It remains unclear how MexB acquires its LTO form. Here by performing functional and cryo-EM structural investigations of MexB at various stages of the assembly process, we unveil that MexB inserted in lipid membrane is not set for active transport because it displays an inactive LTC form with a Closed exit gate. In the tripartite complex, OprM and MexA form a corset-like platform that converts MexB into the active form. Our findings shed new light on the resistance nodulation cell division (RND) cognate partners which act as allosteric factors eliciting the functional drug extrusion. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10371.map.gz emd_10371.map.gz | 96.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10371-v30.xml emd-10371-v30.xml emd-10371.xml emd-10371.xml | 11.5 KB 11.5 KB | Display Display |  EMDB header EMDB header |
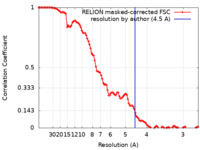
| FSC (resolution estimation) |  emd_10371_fsc.xml emd_10371_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_10371.png emd_10371.png | 145.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10371 http://ftp.pdbj.org/pub/emdb/structures/EMD-10371 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10371 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10371 | HTTPS FTP |
-Related structure data
| Related structure data |  6t7sMC  6ta5C  6ta6C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10371.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10371.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM structure of RND transport MexB trimer from Pseudomonas aeruginosa stabilized in nanodisc | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.36 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : MexB
| Entire | Name: MexB |
|---|---|
| Components |
|
-Supramolecule #1: MexB
| Supramolecule | Name: MexB / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
-Macromolecule #1: MEXB
| Macromolecule | Name: MEXB / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MSKFFIDRPI FAWVIALVIM LAGGLSILSL PVNQYPAIAP PAIAVQVSYP GASAETVQDT VVQVIEQQM NGIDNLRYIS SESNSDGSMT ITVTFEQGTD PDIAQVQVQN KLQLATPLLP Q EVQRQGIR VTKAVKNFLM VVGVVSTDGS MTKEDLSNYI VSNIQDPLSR ...String: MSKFFIDRPI FAWVIALVIM LAGGLSILSL PVNQYPAIAP PAIAVQVSYP GASAETVQDT VVQVIEQQM NGIDNLRYIS SESNSDGSMT ITVTFEQGTD PDIAQVQVQN KLQLATPLLP Q EVQRQGIR VTKAVKNFLM VVGVVSTDGS MTKEDLSNYI VSNIQDPLSR TKGVGDFQVF GS QYSMRIW LDPAKLNSYQ LTPGDVSSAI QAQNVQISSG QLGGLPAVKG QQLNATIIGK TRL QTAEQF ENILLKVNPD GSQVRLKDVA DVGLGGQDYS INAQFNGSPA SGIAIKLATG ANAL DTAKA IRQTIANLEP FMPQGMKVVY PYDTTPVVSA SIHEVVKTLG EAILLVFLVM YLFLQ NFRA TLIPTIAVPV VLLGTFGVLA AFGFSINTLT MFGMVLAIGL LVDDAIVVVE NVERVM AEE GLSPREAARK SMGQIQGALV GIAMVLSAVF LPMAFFGGST GVIYRQFSIT IVSAMAL SV IVALILTPAL CATMLKPIEK GDHGEHKGGF FGWFNRMFLS TTHGYERGVA SILKHRAP Y LLIYVVIVAG MIWMFTRIPT AFLPDEDQGV LFAQVQTPPG SSAERTQVVV DSMREYLLE KESSSVSSVF TVTGFNFAGR GQSSGMAFIM LKPWEERPGG ENSVFELAKR AQMHFFSFKD AMVFAFAPP SVLELGNATG FDLFLQDQAG VGHEVLLQAR NKFLMLAAQN PALQRVRPNG M SDEPQYKL EIDDEKASAL GVSLADINST VSIAWGSSYV NDFIDRGRVK RVYLQGRPDA RM NPDDLSK WYVRNDKGEM VPFNAFATGK WEYGSPKLER YNGVPAMEIL GEPAPGLSSG DAM AAVEEI VKQLPKGVGY SWTGLSYEER LSGSQAPALY ALSLLVVFLC LAALYESWSI PFSV MLVVP LGVIGALLAT SMRGLSNDVF FQVGLLTTIG LSAKNAILIV EFAKELHEQG KGIVE AAIE ACRMRLRPIV MTSLAFILGV VPLAISTGAG SGSQHAIGTG VIGGMVTATV LAIFWV PLF YVAVSTLFKD EASKQQASVE KGQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 42.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)