[English] 日本語
 Yorodumi
Yorodumi- EMDB-10182: Cryo-EM structure of Escherichia coli AcrBZ and DARPin in Saposin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10182 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of Escherichia coli AcrBZ and DARPin in Saposin A-nanodisc with cardiolipin | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RND transporter / efflux pump / drug transport / antibiotic resistance / lipid nanodisc / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationalkane transmembrane transporter activity / alkane transport / enterobactin transport / enterobactin transmembrane transporter activity / xenobiotic detoxification by transmembrane export across the cell outer membrane / periplasmic side of plasma membrane / efflux pump complex / Iron assimilation using enterobactin / bile acid transmembrane transporter activity / xenobiotic detoxification by transmembrane export across the plasma membrane ...alkane transmembrane transporter activity / alkane transport / enterobactin transport / enterobactin transmembrane transporter activity / xenobiotic detoxification by transmembrane export across the cell outer membrane / periplasmic side of plasma membrane / efflux pump complex / Iron assimilation using enterobactin / bile acid transmembrane transporter activity / xenobiotic detoxification by transmembrane export across the plasma membrane / Antimicrobial resistance / bile acid and bile salt transport / Secretion of toxins / efflux transmembrane transporter activity / xenobiotic transmembrane transporter activity / membrane => GO:0016020 / fatty acid transport / xenobiotic transport / cell outer membrane / response to toxic substance / response to xenobiotic stimulus / response to antibiotic / membrane / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Shigella boydii 965-58 (bacteria) Shigella boydii 965-58 (bacteria) | |||||||||
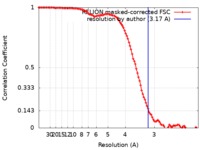
| Method | single particle reconstruction / cryo EM / Resolution: 3.17 Å | |||||||||
 Authors Authors | Szewczak-Harris A / Du D | |||||||||
| Funding support |  United Kingdom, 1 items United Kingdom, 1 items
| |||||||||
 Citation Citation |  Journal: Structure / Year: 2020 Journal: Structure / Year: 2020Title: Interactions of a Bacterial RND Transporter with a Transmembrane Small Protein in a Lipid Environment. Authors: Dijun Du / Arthur Neuberger / Mona Wu Orr / Catherine E Newman / Pin-Chia Hsu / Firdaus Samsudin / Andrzej Szewczak-Harris / Leana M Ramos / Mekdes Debela / Syma Khalid / Gisela Storz / Ben F Luisi /   Abstract: The small protein AcrZ in Escherichia coli interacts with the transmembrane portion of the multidrug efflux pump AcrB and increases resistance of the bacterium to a subset of the antibiotic ...The small protein AcrZ in Escherichia coli interacts with the transmembrane portion of the multidrug efflux pump AcrB and increases resistance of the bacterium to a subset of the antibiotic substrates of that transporter. It is not clear how the physical association of the two proteins selectively changes activity of the pump for defined substrates. Here, we report cryo-EM structures of AcrB and the AcrBZ complex in lipid environments, and comparisons suggest that conformational changes occur in the drug-binding pocket as a result of AcrZ binding. Simulations indicate that cardiolipin preferentially interacts with the AcrBZ complex, due to increased contact surface, and we observe that chloramphenicol sensitivity of bacteria lacking AcrZ is exacerbated when combined with cardiolipin deficiency. Taken together, the data suggest that AcrZ and lipid cooperate to allosterically modulate AcrB activity. This mode of regulation by a small protein and lipid may occur for other membrane proteins. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10182.map.gz emd_10182.map.gz | 96.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10182-v30.xml emd-10182-v30.xml emd-10182.xml emd-10182.xml | 25.8 KB 25.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10182_fsc.xml emd_10182_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_10182.png emd_10182.png | 52.5 KB | ||
| Masks |  emd_10182_msk_1.map emd_10182_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-10182.cif.gz emd-10182.cif.gz | 7.6 KB | ||
| Others |  emd_10182_half_map_1.map.gz emd_10182_half_map_1.map.gz emd_10182_half_map_2.map.gz emd_10182_half_map_2.map.gz | 80.8 MB 80.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10182 http://ftp.pdbj.org/pub/emdb/structures/EMD-10182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10182 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10182 | HTTPS FTP |
-Related structure data
| Related structure data |  6sgrMC  6sgsC  6sgtC  6sguC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10182.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10182.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.088 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10182_msk_1.map emd_10182_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: None
| File | emd_10182_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: None
| File | emd_10182_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ternary complex of AcrBZ and DARPin in Saposin A-nanodisc with ca...
| Entire | Name: Ternary complex of AcrBZ and DARPin in Saposin A-nanodisc with cardiolipin |
|---|---|
| Components |
|
-Supramolecule #1: Ternary complex of AcrBZ and DARPin in Saposin A-nanodisc with ca...
| Supramolecule | Name: Ternary complex of AcrBZ and DARPin in Saposin A-nanodisc with cardiolipin type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|
-Supramolecule #2: Multidrug efflux pump subunit AcrB
| Supramolecule | Name: Multidrug efflux pump subunit AcrB / type: complex / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Supramolecule #3: DARPin
| Supramolecule | Name: DARPin / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Supramolecule #4: Multidrug efflux pump accessory protein AcrZ
| Supramolecule | Name: Multidrug efflux pump accessory protein AcrZ / type: complex / ID: 4 / Parent: 1 / Macromolecule list: #3 |
|---|---|
| Source (natural) | Organism:  Shigella boydii 965-58 (bacteria) Shigella boydii 965-58 (bacteria) |
-Macromolecule #1: Multidrug efflux pump subunit AcrB
| Macromolecule | Name: Multidrug efflux pump subunit AcrB / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 113.66518 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MPNFFIDRPI FAWVIAIIIM LAGGLAILKL PVAQYPTIAP PAVTISASYP GADAKTVQDT VTQVIEQNMN GIDNLMYMSS NSDSTGTVQ ITLTFESGTD ADIAQVQVQN KLQLAMPLLP QEVQQQGVSV EKSSSSFLMV VGVINTDGTM TQEDISDYVA A NMKDAISR ...String: MPNFFIDRPI FAWVIAIIIM LAGGLAILKL PVAQYPTIAP PAVTISASYP GADAKTVQDT VTQVIEQNMN GIDNLMYMSS NSDSTGTVQ ITLTFESGTD ADIAQVQVQN KLQLAMPLLP QEVQQQGVSV EKSSSSFLMV VGVINTDGTM TQEDISDYVA A NMKDAISR TSGVGDVQLF GSQYAMRIWM NPNELNKFQL TPVDVITAIK AQNAQVAAGQ LGGTPPVKGQ QLNASIIAQT RL TSTEEFG KILLKVNQDG SRVLLRDVAK IELGGENYDI IAEFNGQPAS GLGIKLATGA NALDTAAAIR AELAKMEPFF PSG LKIVYP YDTTPFVKIS IHEVVKTLVE AIILVFLVMY LFLQNFRATL IPTIAVPVVL LGTFAVLAAF GFSINTLTMF GMVL AIGLL VDDAIVVVEN VERVMAEEGL PPKEATRKSM GQIQGALVGI AMVLSAVFVP MAFFGGSTGA IYRQFSITIV SAMAL SVLV ALILTPALCA TMLKPIAKGD HGEGKKGFFG WFNRMFEKST HHYTDSVGGI LRSTGRYLVL YLIIVVGMAY LFVRLP SSF LPDEDQGVFM TMVQLPAGAT QERTQKVLNE VTHYYLTKEK NNVESVFAVN GFGFAGRGQN TGIAFVSLKD WADRPGE EN KVEAITMRAT RAFSQIKDAM VFAFNLPAIV ELGTATGFDF ELIDQAGLGH EKLTQARNQL LAEAAKHPDM LTSVRPNG L EDTPQFKIDI DQEKAQALGV SINDINTTLG AAWGGSYVND FIDRGRVKKV YVMSEAKYRM LPDDIGDWYV RAADGQMVP FSAFSSSRWE YGSPRLERYN GLPSMEILGQ AAPGKSTGEA MELMEQLASK LPTGVGYDWT GMSYQERLSG NQAPSLYAIS LIVVFLCLA ALYESWSIPF SVMLVVPLGV IGALLAATFR GLTNDVYFQV GLLTTIGLSA KNAILIVEFA KDLMDKEGKG L IEATLDAV RMRLRPILMT SLAFILGVMP LVISTGAGSG AQNAVGTGVM GGMVTATVLA IFFVPVFFVV VRRRFSRKNE DI EHSHTVD HH UniProtKB: Multidrug efflux pump subunit AcrB |
-Macromolecule #2: DARPin
| Macromolecule | Name: DARPin / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 18.317566 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRGSHHHHHH GSDLGKKLLE AARAGRDDEV RILMANGADV NAADVVGWTP LHLAAYWGHL EIVEVLLKNG ADVNAYDTLG STPLHLAAH FGHLEIVEVL LKNGADVNAK DDNGITPLHL AANRGHLEIV EVLLKYGADV NAQDKFGKTA FDISINNGNE D LAEILQKL N |
-Macromolecule #3: Multidrug efflux pump accessory protein AcrZ
| Macromolecule | Name: Multidrug efflux pump accessory protein AcrZ / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 5.304423 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MLELLKSLVF AVIMVPVVMA IILGLIYGLG EVFNIFSGVG KKDQPGQNH UniProtKB: Multidrug efflux pump accessory protein AcrZ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2.0 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.0038 kPa | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV | |||||||||
| Details | Monodisperse |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 2720 / Average exposure time: 60.0 sec. / Average electron dose: 46.9 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.4 µm / Nominal defocus min: 1.3 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)