[English] 日本語
 Yorodumi
Yorodumi- EMDB-10146: Structure of the RBM2inner region of the Salmonella flagella MS-r... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-10146 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the RBM2inner region of the Salmonella flagella MS-ring protein FliF with 21-fold symmetry applied. | ||||||||||||||||||
 Map data Map data | |||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||
 Keywords Keywords | Flagella / Secretion / Rotor / MS-ring / C-ring / MOTOR PROTEIN | ||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum basal body, MS ring / cytoskeletal motor activity / bacterial-type flagellum-dependent cell motility / plasma membrane Similarity search - Function | ||||||||||||||||||
| Biological species |  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) | ||||||||||||||||||
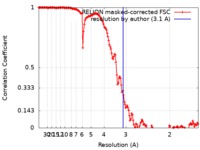
| Method | single particle reconstruction / cryo EM / Resolution: 3.1 Å | ||||||||||||||||||
 Authors Authors | Johnson S / Fong YH | ||||||||||||||||||
| Funding support |  United Kingdom, 5 items United Kingdom, 5 items
| ||||||||||||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2020 Journal: Nat Microbiol / Year: 2020Title: Symmetry mismatch in the MS-ring of the bacterial flagellar rotor explains the structural coordination of secretion and rotation. Authors: Steven Johnson / Yu Hang Fong / Justin C Deme / Emily J Furlong / Lucas Kuhlen / Susan M Lea /  Abstract: The bacterial flagellum is a complex self-assembling nanomachine that confers motility to the cell. Despite great variation across species, all flagella are ultimately constructed from a helical ...The bacterial flagellum is a complex self-assembling nanomachine that confers motility to the cell. Despite great variation across species, all flagella are ultimately constructed from a helical propeller that is attached to a motor embedded in the inner membrane. The motor consists of a series of stator units surrounding a central rotor made up of two ring complexes, the MS-ring and the C-ring. Despite many studies, high-resolution structural information is still lacking for the MS-ring of the rotor, and proposed mismatches in stoichiometry between the two rings have long provided a source of confusion for the field. Here, we present structures of the Salmonella MS-ring, revealing a high level of variation in inter- and intrachain symmetry that provides a structural explanation for the ability of the MS-ring to function as a complex and elegant interface between the two main functions of the flagellum-protein secretion and rotation. | ||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_10146.map.gz emd_10146.map.gz | 5.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-10146-v30.xml emd-10146-v30.xml emd-10146.xml emd-10146.xml | 17.6 KB 17.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_10146_fsc.xml emd_10146_fsc.xml | 15.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_10146.png emd_10146.png | 239.6 KB | ||
| Masks |  emd_10146_msk_1.map emd_10146_msk_1.map | 307.5 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-10146.cif.gz emd-10146.cif.gz | 5.6 KB | ||
| Others |  emd_10146_additional_1.map.gz emd_10146_additional_1.map.gz emd_10146_additional_2.map.gz emd_10146_additional_2.map.gz emd_10146_additional_3.map.gz emd_10146_additional_3.map.gz | 241.6 MB 241.6 MB 238.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-10146 http://ftp.pdbj.org/pub/emdb/structures/EMD-10146 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10146 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-10146 | HTTPS FTP |
-Related structure data
| Related structure data |  6sd2MC  6scnC  6sd1C  6sd3C  6sd4C  6sd5C  6treC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_10146.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_10146.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.822 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_10146_msk_1.map emd_10146_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #3
| File | emd_10146_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #2
| File | emd_10146_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_10146_additional_3.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Homomeric 33mer of Salmonella enterica serovar Typhimurium FliF -...
| Entire | Name: Homomeric 33mer of Salmonella enterica serovar Typhimurium FliF - 21fold symmetric RBM2inner ring |
|---|---|
| Components |
|
-Supramolecule #1: Homomeric 33mer of Salmonella enterica serovar Typhimurium FliF -...
| Supramolecule | Name: Homomeric 33mer of Salmonella enterica serovar Typhimurium FliF - 21fold symmetric RBM2inner ring type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Molecular weight | Theoretical: 2.02 MDa |
-Macromolecule #1: Flagellar M-ring protein
| Macromolecule | Name: Flagellar M-ring protein / type: protein_or_peptide / ID: 1 / Number of copies: 21 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Molecular weight | Theoretical: 61.295645 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSATASTATQ PKPLEWLNRL RANPRIPLIV AGSAAVAIVV AMVLWAKTPD YRTLFSNLSD QDGGAIVAQL TQMNIPYRFA NGSGAIEVP ADKVHELRLR LAQQGLPKGG AVGFELLDQE KFGISQFSEQ VNYQRALEGE LARTIETLGP VKSARVHLAM P KPSLFVRE ...String: MSATASTATQ PKPLEWLNRL RANPRIPLIV AGSAAVAIVV AMVLWAKTPD YRTLFSNLSD QDGGAIVAQL TQMNIPYRFA NGSGAIEVP ADKVHELRLR LAQQGLPKGG AVGFELLDQE KFGISQFSEQ VNYQRALEGE LARTIETLGP VKSARVHLAM P KPSLFVRE QKSPSASVTV TLEPGRALDE GQISAVVHLV SSAVAGLPPG NVTLVDQSGH LLTQSNTSGR DLNDAQLKFA ND VESRIQR RIEAILSPIV GNGNVHAQVT AQLDFANKEQ TEEHYSPNGD ASKATLRSRQ LNISEQVGAG YPGGVPGALS NQP APPNEA PIATPPTNQQ NAQNTPQTST STNSNSAGPR STQRNETSNY EVDRTIRHTK MNVGDIERLS VAVVVNYKTL ADGK PLPLT ADQMKQIEDL TREAMGFSDK RGDTLNVVNS PFSAVDNTGG ELPFWQQQSF IDQLLAAGRW LLVLVVAWIL WRKAV RPQL TRRVEEAKAA QEQAQVRQET EEAVEVRLSK DEQLQQRRAN QRLGAEVMSQ RIREMSDNDP RVVALVIRQW MSNDHE UniProtKB: Flagellar M-ring protein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 48.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-6sd2: |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)