[English] 日本語
 Yorodumi
Yorodumi- EMDB-0940: Cryo-EM structure of the human PAC1 receptor coupled to an engine... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0940 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the human PAC1 receptor coupled to an engineered heterotrimeric G protein | |||||||||
 Map data Map data | None | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | CLASS B GPCR / PACAP / PAC1R / SIGNALING PROTEIN-HORMONE COMPLEX | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of response to reactive oxygen species / pituitary adenylate cyclase activating polypeptide activity / development of primary female sexual characteristics / type 1 vasoactive intestinal polypeptide receptor binding / type 2 vasoactive intestinal polypeptide receptor binding / pituitary adenylate cyclase-activating polypeptide receptor activity / vasoactive intestinal polypeptide receptor activity / positive regulation of growth hormone secretion / positive regulation of chemokine (C-C motif) ligand 5 production / positive regulation of small GTPase mediated signal transduction ...negative regulation of response to reactive oxygen species / pituitary adenylate cyclase activating polypeptide activity / development of primary female sexual characteristics / type 1 vasoactive intestinal polypeptide receptor binding / type 2 vasoactive intestinal polypeptide receptor binding / pituitary adenylate cyclase-activating polypeptide receptor activity / vasoactive intestinal polypeptide receptor activity / positive regulation of growth hormone secretion / positive regulation of chemokine (C-C motif) ligand 5 production / positive regulation of small GTPase mediated signal transduction / neuropeptide hormone activity / regulation of G protein-coupled receptor signaling pathway / NGF-independant TRKA activation / G-protein activation / Activation of the phototransduction cascade / Glucagon-type ligand receptors / Thromboxane signalling through TP receptor / Sensory perception of sweet, bitter, and umami (glutamate) taste / G beta:gamma signalling through PI3Kgamma / G beta:gamma signalling through CDC42 / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / Activation of G protein gated Potassium channels / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / Ca2+ pathway / G alpha (z) signalling events / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / G protein-coupled peptide receptor activity / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / Vasopressin regulates renal water homeostasis via Aquaporins / Adrenaline,noradrenaline inhibits insulin secretion / ADP signalling through P2Y purinoceptor 12 / positive regulation of protein kinase activity / G alpha (q) signalling events / G alpha (i) signalling events / Thrombin signalling through proteinase activated receptors (PARs) / neuropeptide binding / Activation of G protein gated Potassium channels / G-protein activation / G beta:gamma signalling through PI3Kgamma / Prostacyclin signalling through prostacyclin receptor / G beta:gamma signalling through PLC beta / ADP signalling through P2Y purinoceptor 1 / Thromboxane signalling through TP receptor / Presynaptic function of Kainate receptors / G beta:gamma signalling through CDC42 / Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits / insulin secretion / G alpha (12/13) signalling events / Glucagon-type ligand receptors / G beta:gamma signalling through BTK / ADP signalling through P2Y purinoceptor 12 / Adrenaline,noradrenaline inhibits insulin secretion / Cooperation of PDCL (PhLP1) and TRiC/CCT in G-protein beta folding / Ca2+ pathway / G alpha (z) signalling events / Thrombin signalling through proteinase activated receptors (PARs) / Extra-nuclear estrogen signaling / G alpha (s) signalling events / G alpha (q) signalling events / photoreceptor outer segment membrane / spectrin binding / G alpha (i) signalling events / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / peptide hormone receptor binding / High laminar flow shear stress activates signaling by PIEZO1 and PECAM1:CDH5:KDR in endothelial cells / Vasopressin regulates renal water homeostasis via Aquaporins / alkylglycerophosphoethanolamine phosphodiesterase activity / positive regulation of inositol phosphate biosynthetic process / adenylate cyclase-activating G protein-coupled bile acid receptor signaling pathway / adenylate cyclase-activating serotonin receptor signaling pathway / negative regulation of cell cycle / peptide hormone binding / positive regulation of cAMP/PKA signal transduction / regulation of skeletal muscle contraction / adenylate cyclase binding / positive regulation of GTPase activity / PKA activation in glucagon signalling / positive regulation of calcium ion transport into cytosol / hair follicle placode formation / developmental growth / bicellular tight junction / intracellular transport / photoreceptor outer segment / cAMP/PKA signal transduction / D1 dopamine receptor binding / multicellular organismal response to stress / vascular endothelial cell response to laminar fluid shear stress / renal water homeostasis / activation of adenylate cyclase activity / Hedgehog 'off' state / cellular response to acidic pH / adenylate cyclase-activating adrenergic receptor signaling pathway / cardiac muscle cell apoptotic process / photoreceptor inner segment / cellular response to glucagon stimulus / intracellular glucose homeostasis / positive regulation of insulin secretion involved in cellular response to glucose stimulus / adenylate cyclase activator activity / trans-Golgi network membrane / neuropeptide signaling pathway Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.9 Å | |||||||||
 Authors Authors | Kobayashi K / Shihoya W | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2020 Journal: Nat Struct Mol Biol / Year: 2020Title: Cryo-EM structure of the human PAC1 receptor coupled to an engineered heterotrimeric G protein. Authors: Kazuhiro Kobayashi / Wataru Shihoya / Tomohiro Nishizawa / Francois Marie Ngako Kadji / Junken Aoki / Asuka Inoue / Osamu Nureki /  Abstract: Pituitary adenylate cyclase-activating polypeptide (PACAP) is a pleiotropic neuropeptide hormone. The PACAP receptor PAC1R, which belongs to the class B G-protein-coupled receptors (GPCRs), is a drug ...Pituitary adenylate cyclase-activating polypeptide (PACAP) is a pleiotropic neuropeptide hormone. The PACAP receptor PAC1R, which belongs to the class B G-protein-coupled receptors (GPCRs), is a drug target for mental disorders and dry eye syndrome. Here, we present a cryo-EM structure of human PAC1R bound to PACAP and an engineered G heterotrimer. The structure revealed that transmembrane helix TM1 plays an essential role in PACAP recognition. The extracellular domain (ECD) of PAC1R tilts by ~40° compared with that of the glucagon-like peptide-1 receptor (GLP-1R) and thus does not cover the peptide ligand. A functional analysis demonstrated that the PAC1R ECD functions as an affinity trap and is not required for receptor activation, whereas the GLP-1R ECD plays an indispensable role in receptor activation, illuminating the functional diversity of the ECDs in class B GPCRs. Our structural information will facilitate the design and improvement of better PAC1R agonists for clinical applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0940.map.gz emd_0940.map.gz | 4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0940-v30.xml emd-0940-v30.xml emd-0940.xml emd-0940.xml | 19.7 KB 19.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0940_fsc.xml emd_0940_fsc.xml | 6.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_0940.png emd_0940.png | 87.4 KB | ||
| Filedesc metadata |  emd-0940.cif.gz emd-0940.cif.gz | 6.4 KB | ||
| Others |  emd_0940_half_map_1.map.gz emd_0940_half_map_1.map.gz emd_0940_half_map_2.map.gz emd_0940_half_map_2.map.gz | 19.8 MB 19.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0940 http://ftp.pdbj.org/pub/emdb/structures/EMD-0940 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0940 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0940 | HTTPS FTP |
-Related structure data
| Related structure data |  6lpbMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10351 (Title: Cryo-EM structure of the human PAC1 receptor coupled to an engineered heterotrimeric G protein EMPIAR-10351 (Title: Cryo-EM structure of the human PAC1 receptor coupled to an engineered heterotrimeric G proteinData size: 3.5 TB Data #1: Unaligned movies for the human PAC1 receptor coupled to Gs (dimer) [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0940.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0940.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.30406 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: None
| File | emd_0940_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
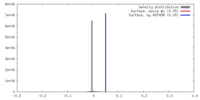
| Density Histograms |
-Half map: None
| File | emd_0940_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | None | ||||||||||||
| Projections & Slices |
| ||||||||||||
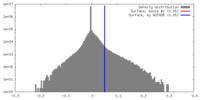
| Density Histograms |
- Sample components
Sample components
-Entire : PAC1R-Gs trimer
| Entire | Name: PAC1R-Gs trimer |
|---|---|
| Components |
|
-Supramolecule #1: PAC1R-Gs trimer
| Supramolecule | Name: PAC1R-Gs trimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Pituitary adenylate cyclase-activating polypeptide
| Macromolecule | Name: Pituitary adenylate cyclase-activating polypeptide / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 4.547336 KDa |
| Sequence | String: HSDGIFTDSY SRYRKQMAVK KYLAAVLGKR YKQRVKNK UniProtKB: Pituitary adenylate cyclase-activating polypeptide |
-Macromolecule #2: Pituitary adenylate cyclase-activating polypeptide type I receptor
| Macromolecule | Name: Pituitary adenylate cyclase-activating polypeptide type I receptor type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 46.67552 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MHSDCIFKKE QAMCLEKIQR ANELMGFNDS SPGCPGMWDN ITCWKPAHVG EMVLVSCPEL FRIFNPDQVW ETETIGESDF GDSNSLDLS DMGVVSRNCT EDGWSEPFPH YFDACGFDEY ESETGDQDYY YLSVKALYTV GYSTSLVTLT TAMVILCRFR K LHCTRNFI ...String: MHSDCIFKKE QAMCLEKIQR ANELMGFNDS SPGCPGMWDN ITCWKPAHVG EMVLVSCPEL FRIFNPDQVW ETETIGESDF GDSNSLDLS DMGVVSRNCT EDGWSEPFPH YFDACGFDEY ESETGDQDYY YLSVKALYTV GYSTSLVTLT TAMVILCRFR K LHCTRNFI HMNLFVSFML RAISVFIKDW ILYAEQDSNH CFISTVECKA VMVFFHYCVV SNYFWLFIEG LYLFTLLVET FF PERRYFY WYTIIGWGTP TVCVTVWATL RLYFDDTGCW DMNDSTALWW VIKGPVVGSI MVNFVLFIGI IVILVQKLQS PDM GGNESS IYLRLARSTL LLIPLFGIHY TVFAFSPENV SKRERLVFEL GLGSFQGFVV AVLYCFLNGE VQAEIKRKWR SENL YFQ UniProtKB: Pituitary adenylate cyclase-activating polypeptide type I receptor |
-Macromolecule #3: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.744371 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHGSL LQSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSRL LVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE LAGHTGYLSC C RFLDDNQI ...String: MHHHHHHGSL LQSELDQLRQ EAEQLKNQIR DARKACADAT LSQITNNIDP VGRIQMRTRR TLRGHLAKIY AMHWGTDSRL LVSASQDGK LIIWDSYTTN KVHAIPLRSS WVMTCAYAPS GNYVACGGLD NICSIYNLKT REGNVRVSRE LAGHTGYLSC C RFLDDNQI VTSSGDTTCA LWDIETGQQT TTFTGHTGDV MSLSLAPDTR LFVSGACDAS AKLWDVREGM CRQTFTGHES DI NAICFFP NGNAFATGSD DATCRLFDLR ADQELMTYSH DNIICGITSV SFSKSGRLLL AGYDDFNCNV WDALKADRAG VLA GHDNRV SCLGVTDDGM AVATGSWDSF LKIWN UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 |
-Macromolecule #4: Guanine nucleotide-binding protein G(s) subunit alpha isoforms sh...
| Macromolecule | Name: Guanine nucleotide-binding protein G(s) subunit alpha isoforms short,Guanine nucleotide-binding protein G(s) subunit alpha isoforms short type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 28.964734 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GNSKTEDQRN EEKAQREANK KIEKQLQKDK QVYRATHRLL LLGADNSGKS TIVKQMRILH GGSGGSGGTS GIFETKFQVD KVNFHMFDV GGQRDERRKW IQCFNDVTAI IFVVDSSDYN RLQEALNLFK SIWNNRWLRT ISVILFLNKQ DLLAEKVLAG K SKIEDYFP ...String: GNSKTEDQRN EEKAQREANK KIEKQLQKDK QVYRATHRLL LLGADNSGKS TIVKQMRILH GGSGGSGGTS GIFETKFQVD KVNFHMFDV GGQRDERRKW IQCFNDVTAI IFVVDSSDYN RLQEALNLFK SIWNNRWLRT ISVILFLNKQ DLLAEKVLAG K SKIEDYFP EFARYTTPED ATPEPGEDPR VTRAKYFIRD EFLRISTASG DGRHYCYPHF TCAVDTENAR RIFNDCRDII QR MHLRQYE LL |
-Macromolecule #5: nanobody Nb35
| Macromolecule | Name: nanobody Nb35 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: unidentified (others) |
| Molecular weight | Theoretical: 15.015728 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MGQVQLQESG GGLVQPGGSL RLSCAASGFT FSNYKMNWVR QAPGKGLEWV SDISQSGASI SYTGSVKGRF TISRDNAKNT LYLQMNSLK PEDTAVYYCA RCPAPFTRDC FDVTSTTYAY RGQGTQVTVS SLHHHHHH |
-Macromolecule #6: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2
| Macromolecule | Name: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 7.547685 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MASNNTASIA QARKLVEQLK MEANIDRIKV SKAAADLMAY CEAHAKEDPL LTPVPASENP FREKKFFS UniProtKB: Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER/RHODIUM / Mesh: 300 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Average electron dose: 64.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)