[English] 日本語
 Yorodumi
Yorodumi- EMDB-0840: Cryo-EM structure of the Drosophila CTP synthase substrate-bound ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0840 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the Drosophila CTP synthase substrate-bound filament | |||||||||
 Map data Map data | the structure of CTPs from drosophila with substrate | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | substrate-bound / filament / LIGASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationInterconversion of nucleotide di- and triphosphates / larval lymph gland hemopoiesis / CTP synthase (glutamine hydrolysing) / CTP synthase activity / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / CTP biosynthetic process / ATP binding / identical protein binding ...Interconversion of nucleotide di- and triphosphates / larval lymph gland hemopoiesis / CTP synthase (glutamine hydrolysing) / CTP synthase activity / cytoophidium / 'de novo' CTP biosynthetic process / pyrimidine nucleobase biosynthetic process / CTP biosynthetic process / ATP binding / identical protein binding / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
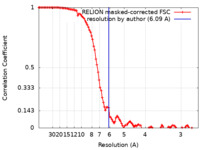
| Method | helical reconstruction / cryo EM / Resolution: 6.09 Å | |||||||||
 Authors Authors | Ji-Long L / Xian Z | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: J Genet Genomics / Year: 2019 Journal: J Genet Genomics / Year: 2019Title: Drosophila CTP synthase can form distinct substrate- and product-bound filaments. Authors: Xian Zhou / Chen-Jun Guo / Huan-Huan Hu / Jiale Zhong / Qianqian Sun / Dandan Liu / Shuang Zhou / Chia Chun Chang / Ji-Long Liu /  Abstract: Intracellular compartmentation is a key strategy for the functioning of a cell. In 2010, several studies revealed that the metabolic enzyme CTP synthase (CTPS) can form filamentous structures termed ...Intracellular compartmentation is a key strategy for the functioning of a cell. In 2010, several studies revealed that the metabolic enzyme CTP synthase (CTPS) can form filamentous structures termed cytoophidia in prokaryotic and eukaryotic cells. However, recent structural studies showed that CTPS only forms inactive product-bound filaments in bacteria while forming active substrate-bound filaments in eukaryotic cells. In this study, using negative staining and cryo-electron microscopy, we demonstrate that Drosophila CTPS, whether in substrate-bound or product-bound form, can form filaments. Our results challenge the previous model and indicate that substrate-bound and product-bound filaments can coexist in the same species. We speculate that the ability to switch between active and inactive cytoophidia in the same cells provides an additional layer of metabolic regulation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0840.map.gz emd_0840.map.gz | 6.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0840-v30.xml emd-0840-v30.xml emd-0840.xml emd-0840.xml | 13.8 KB 13.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0840_fsc.xml emd_0840_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_0840.png emd_0840.png | 30.2 KB | ||
| Masks |  emd_0840_msk_1.map emd_0840_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-0840.cif.gz emd-0840.cif.gz | 5.4 KB | ||
| Others |  emd_0840_half_map_1.map.gz emd_0840_half_map_1.map.gz emd_0840_half_map_2.map.gz emd_0840_half_map_2.map.gz | 6.6 MB 6.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0840 http://ftp.pdbj.org/pub/emdb/structures/EMD-0840 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0840 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0840 | HTTPS FTP |
-Related structure data
| Related structure data |  6l6zMC  0876C  6lfgC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_0840.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0840.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | the structure of CTPs from drosophila with substrate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.34 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_0840_msk_1.map emd_0840_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 2
| File | emd_0840_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map 1
| File | emd_0840_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Drosophila substrate-bound CTP synthase
| Entire | Name: Drosophila substrate-bound CTP synthase |
|---|---|
| Components |
|
-Supramolecule #1: Drosophila substrate-bound CTP synthase
| Supramolecule | Name: Drosophila substrate-bound CTP synthase / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: CTP synthase
| Macromolecule | Name: CTP synthase / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO / EC number: CTP synthase (glutamine hydrolysing) |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 63.11848 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKYILVTGGV ISGVGKGVIA SSFGTLLKSC GLDVTSIKID PYINIDAGTF SPYEHGEVYV LDDGAEVDLD LGNYERFLDV TLHRDNNIT TGKIYKLVIE KERTGEYLGK TVQVVPHITD AIQEWVERVA QTPVQGSSKP QVCIVELGGT IGDIEGMPFV E AFRQFQFR ...String: MKYILVTGGV ISGVGKGVIA SSFGTLLKSC GLDVTSIKID PYINIDAGTF SPYEHGEVYV LDDGAEVDLD LGNYERFLDV TLHRDNNIT TGKIYKLVIE KERTGEYLGK TVQVVPHITD AIQEWVERVA QTPVQGSSKP QVCIVELGGT IGDIEGMPFV E AFRQFQFR VKRENFCLAH VSLVPLPKAT GEPKTKPTQS SVRELRGCGL SPDLIVCRSE KPIGLEVKEK ISNFCHVGPD QV ICIHDLN SIYHVPLLME QNGVIEYLNE RLQLNIDMSK RTKCLQQWRD LARRTETVRR EVCIAVVGKY TKFTDSYASV VKA LQHAAL AVNRKLELVF IESCLLEEET LHSEPSKYHK EWQKLCDSHG ILVPGGFGSR GMEGKIRACQ WARENQKPLL GICL GLQAA VIEFARNKLG LKDANTTEID PNTANALVID MPEHHTGQLG GTMRLGKRIT VFSDGPSVIR QLYGNPKSVQ ERHRH RYEV NPKYVHLLEE QGMRFVGTDV DKTRMEIIEL SGHPYFVATQ YHPEYLSRPL KPSPPFLGLI LASVDRLNQY IQRGCR LS UniProtKB: CTP synthase |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | helical reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 45.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: DARK FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)