+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-0823 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
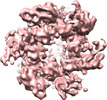
| Title | Cryo-EM map of human MCM8/9 complex | |||||||||
 Map data Map data | Use lower contour value for C terminal display | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
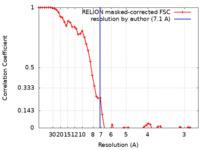
| Method | single particle reconstruction / cryo EM / Resolution: 7.1 Å | |||||||||
 Authors Authors | Li J / Yu D / Liu L / Zeng H / Hu Z / Xu H / Liu B / Liang H / Ouyang Q / Liu Y | |||||||||
 Citation Citation |  Journal: Structure / Year: 2021 Journal: Structure / Year: 2021Title: Structural study of the N-terminal domain of human MCM8/9 complex. Authors: Jun Li / Daqi Yu / Lan Liu / Huanhuan Liang / Qi Ouyang / Yingfang Liu /  Abstract: MCM8/9 is a complex involved in homologous recombination (HR) repair pathway. MCM8/9 dysfunction can cause genome instability and result in primary ovarian insufficiency (POI). However, the mechanism ...MCM8/9 is a complex involved in homologous recombination (HR) repair pathway. MCM8/9 dysfunction can cause genome instability and result in primary ovarian insufficiency (POI). However, the mechanism underlying these effects is largely unknown. Here, we report crystal structures of the N-terminal domains (NTDs) of MCM8 and MCM9, and build a ring-shaped NTD structure based on a 6.6 Å resolution cryoelectron microscopy map. This shows that the MCM8/9 complex forms a 3:3 heterohexamer in an alternating pattern. A positively charged DNA binding channel and a putative ssDNA exit pathway for fork DNA unwinding are revealed. Based on the atomic model, the potential effects of the clinical POI mutants are interpreted. Surprisingly, the zinc-finger motifs are found to be capable of binding an iron atom as well. Overall, our results provide a model for the formation of the MCM8/9 complex and provide a path for further studies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_0823.map.gz emd_0823.map.gz | 3.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-0823-v30.xml emd-0823-v30.xml emd-0823.xml emd-0823.xml | 12.4 KB 12.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_0823_fsc.xml emd_0823_fsc.xml | 5.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_0823.png emd_0823.png | 200.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-0823 http://ftp.pdbj.org/pub/emdb/structures/EMD-0823 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0823 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-0823 | HTTPS FTP |
-Validation report
| Summary document |  emd_0823_validation.pdf.gz emd_0823_validation.pdf.gz | 348.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_0823_full_validation.pdf.gz emd_0823_full_validation.pdf.gz | 348.1 KB | Display | |
| Data in XML |  emd_0823_validation.xml.gz emd_0823_validation.xml.gz | 8.8 KB | Display | |
| Data in CIF |  emd_0823_validation.cif.gz emd_0823_validation.cif.gz | 11.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0823 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0823 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0823 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-0823 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_0823.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_0823.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Use lower contour value for C terminal display | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.37 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Minichromosome Maintenance (MCM) 8/9 Complex
| Entire | Name: Minichromosome Maintenance (MCM) 8/9 Complex |
|---|---|
| Components |
|
-Supramolecule #1: Minichromosome Maintenance (MCM) 8/9 Complex
| Supramolecule | Name: Minichromosome Maintenance (MCM) 8/9 Complex / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Trichoplusia ni (cabbage looper) Trichoplusia ni (cabbage looper) |
| Molecular weight | Theoretical: 500 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | ||||||
|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.8 / Component:
| ||||||
| Grid | Model: C-flat-1.2/1.3 4C / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE | ||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 7420 pixel / Digitization - Dimensions - Height: 7676 pixel / Digitization - Frames/image: 1-40 / Number real images: 3711 / Average exposure time: 10.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated magnification: 36496 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.7 µm / Nominal defocus min: 0.7000000000000001 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)