[English] 日本語
 Yorodumi
Yorodumi- ChemComp-SUE: (1aR,5S,8S,10R,22aR)-5-tert-butyl-N-{(1R,2S)-1-[(cyclopropylsulfo... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 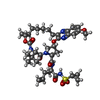 Database: PDB chemical components / ID: SUE Database: PDB chemical components / ID: SUE | ||
|---|---|---|---|
| Name | Name: ( Synonyms: Grazoprevir, MK-5172 Comment | protease inhibitor*YM | |
-Chemical information
| Composition |  | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: SUE / Ideal coordinates details: Corina / Model coordinates PDB-ID: 3SUD | ||||||||||
| History |
| ||||||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  BindingDB / BindingDB /  Brenda Brenda  / /  ChEBI / ChEBI /  ChEMBL / ChEMBL /  CompTox / CompTox /  DrugBank / DrugBank /  PubChem / PubChem /  SureChEMBL SureChEMBL  / /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 12.01 | | CACTVS 3.370 | OpenEye OEToolkits 1.7.6 | |
|---|
-SMILES CANONICAL
| CACTVS 3.370 | | OpenEye OEToolkits 1.7.6 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-PDB entries
Showing all 6 items

PDB-3sud: 
Crystal structure of NS3/4A protease in complex with MK-5172

PDB-3sue: 
Crystal structure of NS3/4A protease variant R155K in complex with MK-5172

PDB-3suf: 
Crystal structure of NS3/4A protease variant D168A in complex with MK-5172

PDB-3sug: 
Crystal structure of NS3/4A protease variant A156T in complex with MK-5172

PDB-6c2m: 
Crystal structure of HCV NS3/4A protease variant Y56H in complex with MK-5172

PDB-6p6q: 
HCV NS3/4A protease domain of genotype 1a3a chimera in complex with grazoprevir
 Movie
Movie Controller
Controller


