[English] 日本語
 Yorodumi
Yorodumi- ChemComp-SAU: 13-methyl[1,3]benzodioxolo[5,6-c][1,3]dioxolo[4,5-i]phenanthridin... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 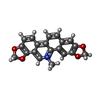 Database: PDB chemical components / ID: SAU Database: PDB chemical components / ID: SAU | ||
|---|---|---|---|
| Name | Name: Synonyms: Sanguinarine Comment | alkaloid*YM | |
-Chemical information
| Composition |  | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: SAU / Ideal coordinates details: Corina / Model coordinates PDB-ID: 3NX5 | ||||||||
| History |
| ||||||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  BindingDB / BindingDB /  Brenda / Brenda /  ChEBI / ChEBI /  ChEMBL / ChEMBL /  ChemicalBook / ChemicalBook /  CompTox / CompTox /  HMDB / HMDB /  KEGG_Ligand / KEGG_Ligand /  Metabolights / Metabolights /  NMRShiftDB / NMRShiftDB /  Nikkaji / Nikkaji /  PubChem / PubChem /  PubChem_TPharma / PubChem_TPharma /  Rhea / Rhea /  SureChEMBL / SureChEMBL /  ZINC / ZINC /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 12.01 | | CACTVS 3.370 | OpenEye OEToolkits 1.7.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.370 | | OpenEye OEToolkits 1.7.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-PDB entries
Showing all 6 items

PDB-3arv: 
Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - complex structure with Sanguinarine

PDB-3as0: 
Crystal Structure Analysis of Chitinase A from Vibrio harveyi with novel inhibitors - W275G mutant complex structure with Sanguinarine

PDB-3nx5: 
The crystal structure of Sanguinarine bound to DNA d(CGTACG)

PDB-5icf: 
Crystal structure of (S)-norcoclaurine 6-O-methyltransferase with S-adenosyl-L-homocysteine and sanguinarine

PDB-5zbz: 
Crystal structure of the DEAD domain of Human eIF4A with sanguinarine

PDB-7c6q: 
Novel natural PPARalpha agonist with a unique binding mode
 Movie
Movie Controller
Controller


