[English] 日本語
 Yorodumi
Yorodumi- SASDF34: Free Nuclear receptor CoRepressor NID (spanning from residue Gln2... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 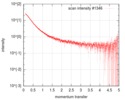 |
|---|---|
 Sample Sample | Free Nuclear receptor CoRepressor NID (spanning from residue Gln2059 to Glu2325)
|
| Function / homology |  Function and homology information Function and homology informationTranscriptional regulation of white adipocyte differentiation / NR1H2 & NR1H3 regulate gene expression to control bile acid homeostasis / NR1H3 & NR1H2 regulate gene expression linked to cholesterol transport and efflux / definitive erythrocyte differentiation / Downregulation of SMAD2/3:SMAD4 transcriptional activity / CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation / Nuclear Receptor transcription pathway / HDACs deacetylate histones / Notch-HLH transcription pathway / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis ...Transcriptional regulation of white adipocyte differentiation / NR1H2 & NR1H3 regulate gene expression to control bile acid homeostasis / NR1H3 & NR1H2 regulate gene expression linked to cholesterol transport and efflux / definitive erythrocyte differentiation / Downregulation of SMAD2/3:SMAD4 transcriptional activity / CD4-positive, CD25-positive, alpha-beta regulatory T cell differentiation / Nuclear Receptor transcription pathway / HDACs deacetylate histones / Notch-HLH transcription pathway / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / Regulation of lipid metabolism by PPARalpha / Cytoprotection by HMOX1 / nuclear retinoic acid receptor binding / thalamus development / regulation of thyroid hormone receptor signaling pathway / nuclear thyroid hormone receptor binding / histone deacetylase regulator activity / locomotor rhythm / cellular response to Thyroglobulin triiodothyronine / regulation of multicellular organism growth / epidermis development / establishment of skin barrier / nuclear retinoid X receptor binding / transcription repressor complex / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / cholesterol homeostasis / circadian regulation of gene expression / transcription corepressor activity / T cell differentiation in thymus / chromatin organization / transcription regulator complex / gene expression / sequence-specific DNA binding / in utero embryonic development / transcription cis-regulatory region binding / protein stabilization / protein domain specific binding / negative regulation of DNA-templated transcription / chromatin binding / regulation of transcription by RNA polymerase II / negative regulation of transcription by RNA polymerase II / DNA binding / nucleoplasm / nucleus / cytosol / cytoplasm Similarity search - Function |
| Biological species | Mouse (Mus musculus) |
 Citation Citation |  Journal: Structure / Year: 2019 Journal: Structure / Year: 2019Title: Interplay of Protein Disorder in Retinoic Acid Receptor Heterodimer and Its Corepressor Regulates Gene Expression. Authors: Tiago N Cordeiro / Nathalie Sibille / Pierre Germain / Philippe Barthe / Abdelhay Boulahtouf / Fréderic Allemand / Rémy Bailly / Valérie Vivat / Christine Ebel / Alessandro Barducci / ...Authors: Tiago N Cordeiro / Nathalie Sibille / Pierre Germain / Philippe Barthe / Abdelhay Boulahtouf / Fréderic Allemand / Rémy Bailly / Valérie Vivat / Christine Ebel / Alessandro Barducci / William Bourguet / Albane le Maire / Pau Bernadó /    Abstract: In its unliganded form, the retinoic acid receptor (RAR) in heterodimer with the retinoid X receptor (RXR) exerts a strong repressive activity facilitated by the recruitment of transcriptional ...In its unliganded form, the retinoic acid receptor (RAR) in heterodimer with the retinoid X receptor (RXR) exerts a strong repressive activity facilitated by the recruitment of transcriptional corepressors in the promoter region of target genes. By integrating complementary structural, biophysical, and computational information, we demonstrate that intrinsic disorder is a required feature for the precise regulation of RAR activity. We show that structural dynamics of RAR and RXR H12 regions is an essential mechanism for RAR regulation. Unexpectedly we found that, while mainly disordered, the corepressor N-CoR presents evolutionary conserved structured regions involved in transient intramolecular contacts. In the presence of RXR/RAR, N-CoR exploits its multivalency to form a cooperative multisite complex that displays equilibrium between different conformational states that can be tuned by cognate ligands and receptor mutations. This equilibrium is key to preserving the repressive basal state while allowing the conversion to a transcriptionally active form. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDF34 SASDF34 |
|---|
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
- Sample
Sample
 Sample Sample | Name: Free Nuclear receptor CoRepressor NID (spanning from residue Gln2059 to Glu2325) Specimen concentration: 2 mg/ml |
|---|---|
| Buffer | Name: 50mM Tris HCl, 150mM NaCl, 2mM TCEP. / pH: 7.5 |
| Entity #1515 | Name: N-CoR-NID / Type: protein Description: Nuclear receptor CoRepressor 1; Nuclear Receptor Interaction Domain (NID) Formula weight: 29.139 / Num. of mol.: 1 / Source: Mouse (Mus musculus) / References: UniProt: Q60974 Sequence: GPHMQVPRTH RLITLADHIC QIITQDFARN QVPSQASTST FQTSPSALSS TPVRTKTSSR YSPESQSQTV LHPRPGPRVS PENLVDKSRG SRPGKSPERS HIPSEPYEPI SPPQGPAVHE KQDSMLLLSQ RGVDPAEQRS DSRSPGSISY LPSFFTKLES TSPMVKSKKQ ...Sequence: GPHMQVPRTH RLITLADHIC QIITQDFARN QVPSQASTST FQTSPSALSS TPVRTKTSSR YSPESQSQTV LHPRPGPRVS PENLVDKSRG SRPGKSPERS HIPSEPYEPI SPPQGPAVHE KQDSMLLLSQ RGVDPAEQRS DSRSPGSISY LPSFFTKLES TSPMVKSKKQ EIFRKLNSSG GGDSDMAAAQ PGTEIFNLPA VTTSGAVSSR SHSFADPASN LGLEDIIRKA LMGSFDDKVE DHGVVMSHPV GIMPGSASTS VVTSSEARRD E |
-Experimental information
| Beam | Instrument name: ESRF BM29 / City: Grenoble / 国: France  / Type of source: X-ray synchrotron / Wavelength: 0.099 Å / Dist. spec. to detc.: 2.9 mm / Type of source: X-ray synchrotron / Wavelength: 0.099 Å / Dist. spec. to detc.: 2.9 mm | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 1M / Type: Dectris / Pixsize x: 172 mm | ||||||||||||||||||||||||||||||||||||||||||
| Scan | Measurement date: Jun 20, 2016 / Storage temperature: 20 °C / Cell temperature: 20 °C / Exposure time: 1 sec. / Number of frames: 15 / Unit: 1/nm /
| ||||||||||||||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| ||||||||||||||||||||||||||||||||||||||||||
| Result | Comments: Several datasets were collected at multiple concentrations ranging between 1.3 and 2.0 mg/ml. Only the data of 2mg/ml is shown.
|
 Movie
Movie Controller
Controller