+ Open data
Open data
- Basic information
Basic information
| Entry |  |
|---|---|
 Sample Sample | Ribonuclease E from Yersinia pestis
|
| Function / homology |  Function and homology information Function and homology informationribonuclease E / ribonuclease E activity / tRNA processing / mRNA catabolic process / RNA nuclease activity / RNA endonuclease activity / cytoplasmic side of plasma membrane / rRNA processing / tRNA binding / rRNA binding ...ribonuclease E / ribonuclease E activity / tRNA processing / mRNA catabolic process / RNA nuclease activity / RNA endonuclease activity / cytoplasmic side of plasma membrane / rRNA processing / tRNA binding / rRNA binding / magnesium ion binding / zinc ion binding / cytoplasm Similarity search - Function |
| Biological species |  |
 Citation Citation |  Date: 2019 Dec Date: 2019 DecTitle: A structural and biochemical comparison of Ribonuclease E homologues from pathogenic bacteria highlights species-specific properties Authors: Mardle C / Shakespeare T / Butt L / Goddard L / Gowers D / Atkins H / Vincent H |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDE62 SASDE62 |
|---|
-Related structure data
| Related structure data | C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
- Sample
Sample
 Sample Sample | Name: Ribonuclease E from Yersinia pestis / Specimen concentration: 6.33 mg/ml |
|---|---|
| Buffer | Name: 10 mM DTT, 10 mM MgCl2, 0.5 M NaCl, 20 mM Tris / pH: 8 |
| Entity #1204 | Name: RNase E / Type: protein / Description: Endoribonuclease E / Formula weight: 61.892 / Num. of mol.: 4 / Source: Yersinia pestis / References: UniProt: Q74TC3 Sequence: HHHHHHHHHH SGSSGHIEGR HMKRMLINAT QQEELRVALV DGQRLYDLDI ESPGHEQKKA NIYKGKITRI EPSLEAAFVD YGAERHGFLP LKEISREYFP NNYSSHGRPN IKDVLREGQE VIVQVDKEER GNKGAALTTF ISLAGSYLVL MPNNPRAGGI SRRIEGDDRT ...Sequence: HHHHHHHHHH SGSSGHIEGR HMKRMLINAT QQEELRVALV DGQRLYDLDI ESPGHEQKKA NIYKGKITRI EPSLEAAFVD YGAERHGFLP LKEISREYFP NNYSSHGRPN IKDVLREGQE VIVQVDKEER GNKGAALTTF ISLAGSYLVL MPNNPRAGGI SRRIEGDDRT ELKEALSSLQ LPDGMGLIVR TAGVGKSADA LQWDLSFRLK HWDAIKKAAE GRPAPFLIHQ ESNVIVRAFR DYLRPDIGEI LIDNPKVLEL AKEHIAALGR PDFSSKIKLY SGEIPLFSHY QIESQIESAF QREVRLPSGG SIVIDTTEAL TAIDINSARA TRGGDIEETA FNTNLEAADE IARQLRLRDL GGLIVIDFID MTPVRHQREV ENRLRDAVRQ DRARIQIGRI SRFGLLEMSR QRLSPSLGES SHHVCPRCSG TGTVRDNESL SLSILRLIEE EALKENTHEV HAIVPVQIAS YLLNEKRESV NAIEKRQGGV RAVIVPNDQM QTPHYSVLRV RKGEEVPSLS YLLPQLHEAE MAQPLEEATI ERKRPEQPAL |
-Experimental information
| Beam | Instrument name: Diamond Light Source B21 / City: Oxfordshire / 国: UK  / Shape: 1 x 5 mm / Type of source: X-ray synchrotron / Wavelength: 0.1 Å / Dist. spec. to detc.: 4.014 mm / Shape: 1 x 5 mm / Type of source: X-ray synchrotron / Wavelength: 0.1 Å / Dist. spec. to detc.: 4.014 mm | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 2M | ||||||||||||||||||||||||||||||
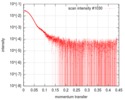
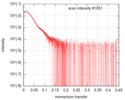
| Scan |
| ||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| ||||||||||||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller