+ Open data
Open data
- Basic information
Basic information
| Entry |  |
|---|---|
 Sample Sample | Candida antarctica lipase B - with guanidine-HCl unfolding series
|
| Function / homology | : / triacylglycerol lipase / triacylglycerol lipase activity / lipid catabolic process / Alpha/Beta hydrolase fold / Lipase B Function and homology information Function and homology information |
| Biological species | Pseudozyma antarctica (Candida antarctica) |
 Citation Citation |  Journal: Biophys J / Year: 2018 Journal: Biophys J / Year: 2018Title: Machine Learning Methods for X-Ray Scattering Data Analysis from Biomacromolecular Solutions. Authors: Daniel Franke / Cy M Jeffries / Dmitri I Svergun /  Abstract: Small-angle x-ray scattering (SAXS) of biological macromolecules in solutions is a widely employed method in structural biology. SAXS patterns include information about the overall shape and low- ...Small-angle x-ray scattering (SAXS) of biological macromolecules in solutions is a widely employed method in structural biology. SAXS patterns include information about the overall shape and low-resolution structure of dissolved particles. Here, we describe how to transform experimental SAXS patterns to feature vectors and how a simple k-nearest neighbor approach is able to retrieve information on overall particle shape and maximal diameter (D) as well as molecular mass directly from experimental scattering data. Based on this transformation, we develop a rapid multiclass shape-classification ranging from compact, extended, and flat categories to hollow and random-chain-like objects. This classification may be employed, e.g., as a decision block in automated data analysis pipelines. Further, we map protein structures from the Protein Data Bank into the classification space and, in a second step, use this mapping as a data source to obtain accurate estimates for the structural parameters (D, molecular mass) of the macromolecule under study based on the experimental scattering pattern alone, without inverse Fourier transform for D. All methods presented are implemented in a Fortran binary DATCLASS, part of the ATSAS data analysis suite, available on Linux, Mac, and Windows and free for academic use. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
- Downloads & links
Downloads & links
-Models
- Sample
Sample
 Sample Sample | Name: Candida antarctica lipase B - with guanidine-HCl unfolding series Specimen concentration: 4.65 mg/ml |
|---|---|
| Buffer | Name: 100 mM NaCl, 20 mM Na2HPO4 / pH: 6 |
| Entity #968 | Name: lipase B / Type: protein / Description: Lipase B from Pseudozyma antarctica / Formula weight: 33.022 / Source: Pseudozyma antarctica (Candida antarctica) / References: UniProt: P41365 Sequence: LPSGSDPAFS QPKSVLDAGL TCQGASPSSV SKPILLVPGT GTTGPQSFDS NWIPLSTQLG YTPCWISPPP FMLNDTQVNT EYMVNAITAL YAGSGNNKLP VLTWSQGGLV AQWGLTFFPS IRSKVDRLMA FAPDYKGTVL AGPLDALAVS APSVWQQTTG SALTTALRNA ...Sequence: LPSGSDPAFS QPKSVLDAGL TCQGASPSSV SKPILLVPGT GTTGPQSFDS NWIPLSTQLG YTPCWISPPP FMLNDTQVNT EYMVNAITAL YAGSGNNKLP VLTWSQGGLV AQWGLTFFPS IRSKVDRLMA FAPDYKGTVL AGPLDALAVS APSVWQQTTG SALTTALRNA GGLTQIVPTT NLYSATDEIV QPQVSNSPLD SSYLFNGKNV QAQAVCGPLF VIDHAGSLTS QFSYVVGRSA LRSTTGQARS ADYGITDCNP LPANDLTPEQ KVAAAALLAP AAAAIVAGPK QNCEPDLMPY ARPFAVGKRT CSGIVTP |
-Experimental information
| Beam | Instrument name: PETRA III EMBL P12 / City: Hamburg / 国: Germany  / Type of source: X-ray synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 3.1 mm / Type of source: X-ray synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 3.1 mm | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 2M | |||||||||||||||||||||
| Scan | Measurement date: Jul 29, 2013 / Storage temperature: 10 °C / Cell temperature: 10 °C / Exposure time: 0.03 sec. / Number of frames: 1 / Unit: 1/nm /
| |||||||||||||||||||||
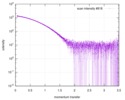
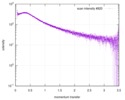
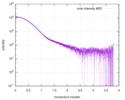
| Result | Type of curve: single_conc Comments: In addition to the SAXS profile displayed above, a lipase B unfolding series spanning 0 M to 6 M guanidine hydrochloride is included in the full entry zip archive.
|
 Movie
Movie Controller
Controller



 SASDDJ3
SASDDJ3