 Yorodumi
Yorodumi+ Open data
Open data
- Basic information
Basic information
| Entry | 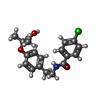 Database: PDB chemical components / ID: PEM Database: PDB chemical components / ID: PEM |
|---|---|
| Name | Name: |
-Chemical information
| Composition |  | ||||
|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: PEM / Ideal coordinates details: OpenEye/OEToolkits V1.4.2 / Model coordinates PDB-ID: 1IWH | ||||
| History |
| ||||
 External links External links |  UniChem / UniChem /  ChemSpider / ChemSpider /  BindingDB BindingDB                                            / /  Brenda / Brenda /  ChEMBL / ChEMBL /  ChemicalBook / ChemicalBook /  CompTox / CompTox /  DrugBank / DrugBank /  GtoPharmacology / GtoPharmacology /  HMDB / HMDB /  Nikkaji / Nikkaji /  PubChem / PubChem /  PubChem_TPharma / PubChem_TPharma /  SureChEMBL / SureChEMBL /  ZINC / ZINC /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 10.04 | | CACTVS 3.341 | OpenEye OEToolkits 1.5.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.341 | | OpenEye OEToolkits 1.5.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 10.04 | | OpenEye OEToolkits 1.5.0 | |
|---|
-PDB entries
Showing all 6 items

PDB-1iwh: 
Crystal Structure of Horse Carbonmonoxyhemoglobin-Bezafibrate Complex at 1.55A Resolution: A Novel Allosteric Binding Site in R-State Hemoglobin

PDB-5x2s: 
Direct Observation of Conformational Population Shifts in Hemoglobin: Crystal Structure of Half-Liganded Hemoglobin after Adding 4 mM bezafibrate pH 6.5.

PDB-5x2t: 
Direct Observation of Conformational Population Shifts in Hemoglobin: Crystal Structure of Half-Liganded Hemoglobin after Adding 4 mM bezafibrate pH 7.2.

PDB-7bpz: 
X-ray structure of human PPARalpha ligand binding domain-bezafibrate-SRC1 coactivator peptide co-crystals obtained by soaking

PDB-7wgl: 
X-ray structure of human PPAR delta ligand binding domain-bezafibrate co-crystals obtained by co-crystallization

PDB-7wgo: 
X-ray structure of human PPAR gamma ligand binding domain-bezafibrate co-rystals obtained by co-crystallization
 Movie
Movie Controller
Controller


