+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8whb | ||||||
|---|---|---|---|---|---|---|---|
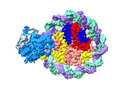
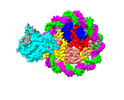
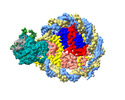
| Title | Structure of nucleosome core particle of Arabidopsis thaliana | ||||||
 Components Components |
| ||||||
 Keywords Keywords | GENE REGULATION / nucleosome / chromatin / histone / protein dna interaction / nucleoprotein | ||||||
| Function / homology |  Function and homology information Function and homology informationDNA-mediated transformation / chloroplast thylakoid / chromocenter / response to water deprivation / plasmodesma / thylakoid / plant-type vacuole / plastid / chloroplast stroma / chloroplast ...DNA-mediated transformation / chloroplast thylakoid / chromocenter / response to water deprivation / plasmodesma / thylakoid / plant-type vacuole / plastid / chloroplast stroma / chloroplast / response to bacterium / response to wounding / structural constituent of chromatin / nucleosome / peroxisome / protein heterodimerization activity / nucleolus / DNA binding / extracellular region / nucleus / plasma membrane / cytosol Similarity search - Function | ||||||
| Biological species |  synthetic construct (others) | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 3.17 Å | ||||||
 Authors Authors | Liu, Y. / Zhang, Z. / Du, J. | ||||||
| Funding support |  China, 1items China, 1items
| ||||||
 Citation Citation |  Journal: Nat Plants / Year: 2024 Journal: Nat Plants / Year: 2024Title: Molecular basis of chromatin remodelling by DDM1 involved in plant DNA methylation. Authors: Yue Liu / Zhihui Zhang / Hongmiao Hu / Wei Chen / Fan Zhang / Qian Wang / Changshi Wang / Kaige Yan / Jiamu Du /   Abstract: Eukaryotic gene regulation occurs at the chromatin level, which requires changing the chromatin structure by a group of ATP-dependent DNA translocases-namely, the chromatin remodellers. In plants, ...Eukaryotic gene regulation occurs at the chromatin level, which requires changing the chromatin structure by a group of ATP-dependent DNA translocases-namely, the chromatin remodellers. In plants, chromatin remodellers function in various biological processes and possess both conserved and plant-specific components. DECREASE IN DNA METHYLATION 1 (DDM1) is a plant chromatin remodeller that plays a key role in the maintenance DNA methylation. Here we determined the structures of Arabidopsis DDM1 in complex with nucleosome in ADP-BeF-bound, ADP-bound and nucleotide-free conformations. We show that DDM1 specifically recognizes the H4 tail and nucleosomal DNA. The conformational differences between ADP-BeF-bound, ADP-bound and nucleotide-free DDM1 suggest a chromatin remodelling cycle coupled to ATP binding, hydrolysis and ADP release. This, in turn, triggers conformational changes in the DDM1-bound nucleosomal DNA, which alters the nucleosome structure and promotes DNA sliding. Together, our data reveal the molecular basis of chromatin remodelling by DDM1. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8whb.cif.gz 8whb.cif.gz | 267.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8whb.ent.gz pdb8whb.ent.gz | 198.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8whb.json.gz 8whb.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  8whb_validation.pdf.gz 8whb_validation.pdf.gz | 1 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  8whb_full_validation.pdf.gz 8whb_full_validation.pdf.gz | 1 MB | Display | |
| Data in XML |  8whb_validation.xml.gz 8whb_validation.xml.gz | 38.8 KB | Display | |
| Data in CIF |  8whb_validation.cif.gz 8whb_validation.cif.gz | 60 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/wh/8whb https://data.pdbj.org/pub/pdb/validation_reports/wh/8whb ftp://data.pdbj.org/pub/pdb/validation_reports/wh/8whb ftp://data.pdbj.org/pub/pdb/validation_reports/wh/8whb | HTTPS FTP |
-Related structure data
| Related structure data |  37538MC  8wh5C  8wh8C  8wh9C  8whaC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
-Protein , 4 types, 8 molecules AEBFCGDH
| #1: Protein | Mass: 15300.968 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: HTR2, At1g09200, T12M4.9, HTR3, At3g27360, K1G2.8, HTR13, At5g10390, F12B17_260, HTR9, At5g10400, F12B17_250, HTR1, At5g65360, MNA5.9 Production host:  #2: Protein | Mass: 11436.467 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Gene: At1g07660, F24B9.25, At1g07820, F24B9.8, At2g28740, F8N16.2, T11P11.4, At3g45930, F16L2_140, At3g46320, F18L15.40, At3g53730, F5K20_30, At5g59690, MTH12.10, At5g59970, MMN10.22 Production host:  #3: Protein | Mass: 13680.854 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #4: Protein | Mass: 16474.459 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|
-DNA chain , 2 types, 2 molecules IJ
| #5: DNA chain | Mass: 45123.758 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  |
|---|---|
| #6: DNA chain | Mass: 45626.043 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Structure of the nucleosome core particle of Arabidopsis thaliana Type: COMPLEX / Entity ID: all / Source: RECOMBINANT |
|---|---|
| Molecular weight | Value: 0.20 MDa / Experimental value: NO |
| Source (natural) | Organism:  |
| Source (recombinant) | Organism:  |
| Buffer solution | pH: 8 |
| Specimen | Conc.: 2.5 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 277.15 K |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal defocus max: 2500 nm / Nominal defocus min: 1000 nm / Cs: 2.7 mm |
| Specimen holder | Cryogen: NITROGEN / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Image recording | Electron dose: 50 e/Å2 / Detector mode: SUPER-RESOLUTION / Film or detector model: GATAN K3 (6k x 4k) / Num. of real images: 6776 |
| Image scans | Movie frames/image: 32 / Used frames/image: 1-32 |
- Processing
Processing
| EM software | Name: RELION / Version: 3.1 / Category: 3D reconstruction | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 3898721 | ||||||||||||||||||||||||
| 3D reconstruction | Resolution: 3.17 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 261707 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller




 PDBj
PDBj






































