+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 8k20 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of KEOPS complex from Arabidopsis thaliana | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSFERASE / KEOPS complex / tRNA t6A synthase / RNA-binding protein | ||||||
| Function / homology |  Function and homology information Function and homology informationN(6)-L-threonylcarbamoyladenine synthase activity / EKC/KEOPS complex / tRNA threonylcarbamoyladenosine modification / tRNA processing / non-specific serine/threonine protein kinase / phosphorylation / protein serine/threonine kinase activity / ATP binding / nucleus / metal ion binding / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 3.7 Å | ||||||
 Authors Authors | Zheng, X.X. / Zhu, L. / Duan, L. / Zhang, W.H. | ||||||
| Funding support |  China, 1items China, 1items
| ||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2024 Journal: Nucleic Acids Res / Year: 2024Title: Molecular basis of A. thaliana KEOPS complex in biosynthesizing tRNA t6A. Authors: Xinxing Zheng / Chenchen Su / Lei Duan / Mengqi Jin / Yongtao Sun / Li Zhu / Wenhua Zhang /  Abstract: In archaea and eukaryotes, the evolutionarily conserved KEOPS is composed of four core subunits-Kae1, Bud32, Cgi121 and Pcc1, and a fifth Gon7/Pcc2 that is found in fungi and metazoa. KEOPS ...In archaea and eukaryotes, the evolutionarily conserved KEOPS is composed of four core subunits-Kae1, Bud32, Cgi121 and Pcc1, and a fifth Gon7/Pcc2 that is found in fungi and metazoa. KEOPS cooperates with Sua5/YRDC to catalyze the biosynthesis of tRNA N6-threonylcarbamoyladenosine (t6A), an essential modification needed for fitness of cellular organisms. Biochemical and structural characterizations of KEOPSs from archaea, yeast and humans have determined a t6A-catalytic role for Kae1 and auxiliary roles for other subunits. However, the precise molecular workings of KEOPSs still remain poorly understood. Here, we investigated the biochemical functions of A. thaliana KEOPS and determined a cryo-EM structure of A. thaliana KEOPS dimer. We show that A. thaliana KEOPS is composed of KAE1, BUD32, CGI121 and PCC1, which adopts a conserved overall arrangement. PCC1 dimerization leads to a KEOPS dimer that is needed for an active t6A-catalytic KEOPS-tRNA assembly. BUD32 participates in direct binding of tRNA to KEOPS and modulates the t6A-catalytic activity of KEOPS via its C-terminal tail and ATP to ADP hydrolysis. CGI121 promotes the binding of tRNA to KEOPS and potentiates the t6A-catalytic activity of KEOPS. These data and findings provide insights into mechanistic understanding of KEOPS machineries. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  8k20.cif.gz 8k20.cif.gz | 217.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb8k20.ent.gz pdb8k20.ent.gz | 172.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  8k20.json.gz 8k20.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  8k20_validation.pdf.gz 8k20_validation.pdf.gz | 1.2 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  8k20_full_validation.pdf.gz 8k20_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  8k20_validation.xml.gz 8k20_validation.xml.gz | 58.3 KB | Display | |
| Data in CIF |  8k20_validation.cif.gz 8k20_validation.cif.gz | 85.2 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/k2/8k20 https://data.pdbj.org/pub/pdb/validation_reports/k2/8k20 ftp://data.pdbj.org/pub/pdb/validation_reports/k2/8k20 ftp://data.pdbj.org/pub/pdb/validation_reports/k2/8k20 | HTTPS FTP |
-Related structure data
| Related structure data | 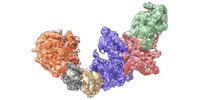 36808MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 38813.570 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   References: UniProt: O49653, N6-L-threonylcarbamoyladenine synthase #2: Protein | | Mass: 25170.117 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   References: UniProt: Q94K14, non-specific serine/threonine protein kinase #3: Protein | | Mass: 19678.484 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #4: Protein | Mass: 11592.994 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #5: Chemical | ChemComp-FE / | Has ligand of interest | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: KEOPS complex from Arabidopsis thaliana / Type: COMPLEX Details: The sample contains KAE1, BUD32, CGI121 and PCC1, which form a KEOPS complex. Entity ID: #1-#4 / Source: RECOMBINANT |
|---|---|
| Molecular weight | Experimental value: NO |
| Source (natural) | Organism:  |
| Source (recombinant) | Organism:  |
| Buffer solution | pH: 7.5 Details: 300 mM NaCl, 20 mM Tris-HCl pH 7.5,5 mM 2-mercaptoethanol |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES Details: The sample was freshly purified by size exclusion chromatography. |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: OTHER FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: OTHER |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 105000 X / Nominal defocus max: 2700 nm / Nominal defocus min: 900 nm / Cs: 0.27 mm / Alignment procedure: COMA FREE |
| Specimen holder | Cryogen: NITROGEN / Temperature (max): 89.8 K / Temperature (min): 77.1 K |
| Image recording | Average exposure time: 3.6 sec. / Electron dose: 64 e/Å2 / Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) |
| Image scans | Width: 5760 / Height: 4092 |
- Processing
Processing
| EM software | Name: PHENIX / Version: 1.19.2_4158: / Category: model refinement | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
| 3D reconstruction | Resolution: 3.7 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 232232 / Symmetry type: POINT | ||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller



 PDBj
PDBj

