+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 7zs9 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
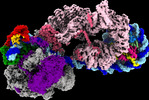
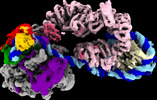
| タイトル | Yeast RNA polymerase II transcription pre-initiation complex with the +1 nucleosome (complex A) | |||||||||
 要素 要素 |
| |||||||||
 キーワード キーワード | TRANSCRIPTION / PIC / nucleosome / TF | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報regulation of mitotic recombination / RNA polymerase II promoter clearance / RNA polymerase II complex recruiting activity / TFIIA-class transcription factor complex binding / RNA polymerase III transcription regulatory region sequence-specific DNA binding / transcription factor TFIIIB complex / RNA polymerase III preinitiation complex assembly / phosphatidylinositol-5-phosphate binding / positive regulation of mitotic recombination / regulation of transcription by RNA polymerase III ...regulation of mitotic recombination / RNA polymerase II promoter clearance / RNA polymerase II complex recruiting activity / TFIIA-class transcription factor complex binding / RNA polymerase III transcription regulatory region sequence-specific DNA binding / transcription factor TFIIIB complex / RNA polymerase III preinitiation complex assembly / phosphatidylinositol-5-phosphate binding / positive regulation of mitotic recombination / regulation of transcription by RNA polymerase III / RNA polymerase I general transcription initiation factor binding / nucleotide-excision repair factor 3 complex / transcription factor TFIIE complex / DNA translocase activity / nucleotide-excision repair, preincision complex assembly / transcription factor TFIIK complex / transcription open complex formation at RNA polymerase II promoter / TFIIF-class transcription factor complex binding / transcriptional start site selection at RNA polymerase II promoter / RNA polymerase II core complex assembly / RPB4-RPB7 complex / positive regulation of transcription regulatory region DNA binding / transcription factor TFIIF complex / phosphatidylinositol-3-phosphate binding / transcription factor TFIIA complex / RNA polymerase I preinitiation complex assembly / transcription factor TFIIH core complex / transcription factor TFIIH holo complex / cyclin-dependent protein serine/threonine kinase activator activity / DNA 3'-5' helicase / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / DNA duplex unwinding / transcription preinitiation complex / DNA binding, bending / RNA Polymerase I Transcription Initiation / Processing of Capped Intron-Containing Pre-mRNA / RNA Polymerase III Transcription Initiation From Type 2 Promoter / poly(A)+ mRNA export from nucleus / RNA Pol II CTD phosphorylation and interaction with CE / Formation of the Early Elongation Complex / mRNA Capping / RNA polymerase II transcribes snRNA genes / TP53 Regulates Transcription of DNA Repair Genes / termination of RNA polymerase II transcription / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / transcription factor TFIID complex / RNA-templated transcription / termination of RNA polymerase III transcription / RNA Polymerase II Pre-transcription Events / RNA polymerase II general transcription initiation factor activity / Formation of TC-NER Pre-Incision Complex / termination of RNA polymerase I transcription / RNA Polymerase I Promoter Escape / transcription initiation at RNA polymerase III promoter / RNA polymerase II complex binding / protein phosphatase activator activity / nucleolar large rRNA transcription by RNA polymerase I / 3'-5' DNA helicase activity / maintenance of transcriptional fidelity during transcription elongation by RNA polymerase II / Gap-filling DNA repair synthesis and ligation in TC-NER / transcription initiation at RNA polymerase I promoter / transcription by RNA polymerase I / nuclear-transcribed mRNA catabolic process / Estrogen-dependent gene expression / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / ATPase activator activity / transcription by RNA polymerase III / acetyltransferase activity / Dual incision in TC-NER / transcription elongation by RNA polymerase I / RNA polymerase II activity / positive regulation of transcription initiation by RNA polymerase II / RNA polymerase II core promoter sequence-specific DNA binding / tRNA transcription by RNA polymerase III / RNA polymerase I complex / RNA polymerase III complex / positive regulation of translational initiation / ATP-dependent activity, acting on DNA / transcription-coupled nucleotide-excision repair / RNA polymerase I activity / RNA polymerase II, core complex / translesion synthesis / RNA polymerase II preinitiation complex assembly / DNA helicase activity / translation initiation factor binding / TBP-class protein binding / DNA-templated transcription initiation / transcription elongation by RNA polymerase II / nucleotide-excision repair / transcription initiation at RNA polymerase II promoter / positive regulation of transcription elongation by RNA polymerase II / P-body / ribonucleoside binding / DNA-directed 5'-3' RNA polymerase activity / DNA-directed RNA polymerase / cytoplasmic stress granule / mRNA processing 類似検索 - 分子機能 | |||||||||
| 生物種 |  synthetic construct (人工物) | |||||||||
| 手法 | 電子顕微鏡法 / 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 3.1 Å | |||||||||
 データ登録者 データ登録者 | Wang, H. / Cramer, P. | |||||||||
| 資金援助 | European Union,  ドイツ, 2件 ドイツ, 2件
| |||||||||
 引用 引用 |  ジャーナル: Nat Struct Mol Biol / 年: 2023 ジャーナル: Nat Struct Mol Biol / 年: 2023タイトル: Structures of transcription preinitiation complex engaged with the +1 nucleosome. 著者: Haibo Wang / Sandra Schilbach / Momchil Ninov / Henning Urlaub / Patrick Cramer /   要旨: The preinitiation complex (PIC) assembles on promoters of protein-coding genes to position RNA polymerase II (Pol II) for transcription initiation. Previous structural studies revealed the PIC on ...The preinitiation complex (PIC) assembles on promoters of protein-coding genes to position RNA polymerase II (Pol II) for transcription initiation. Previous structural studies revealed the PIC on different promoters, but did not address how the PIC assembles within chromatin. In the yeast Saccharomyces cerevisiae, PIC assembly occurs adjacent to the +1 nucleosome that is located downstream of the core promoter. Here we present cryo-EM structures of the yeast PIC bound to promoter DNA and the +1 nucleosome located at three different positions. The general transcription factor TFIIH engages with the incoming downstream nucleosome and its translocase subunit Ssl2 (XPB in human TFIIH) drives the rotation of the +1 nucleosome leading to partial detachment of nucleosomal DNA and intimate interactions between TFIIH and the nucleosome. The structures provide insights into how transcription initiation can be influenced by the +1 nucleosome and may explain why the transcription start site is often located roughly 60 base pairs upstream of the dyad of the +1 nucleosome in yeast. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  7zs9.cif.gz 7zs9.cif.gz | 1.9 MB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb7zs9.ent.gz pdb7zs9.ent.gz | 1.5 MB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  7zs9.json.gz 7zs9.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/zs/7zs9 https://data.pdbj.org/pub/pdb/validation_reports/zs/7zs9 ftp://data.pdbj.org/pub/pdb/validation_reports/zs/7zs9 ftp://data.pdbj.org/pub/pdb/validation_reports/zs/7zs9 | HTTPS FTP |
|---|
-関連構造データ
| 関連構造データ | 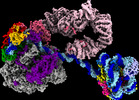 14927MC  7zsaC  7zsbC M: このデータのモデリングに利用したマップデータ C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 
|
|---|---|
| 1 |
|
- 要素
要素
+DNA-directed RNA polymerase II subunit ... , 7種, 7分子 ABCDGIK
+DNA-directed RNA polymerases I, II, and III subunit ... , 5種, 5分子 EFHJL
+タンパク質 , 7種, 11分子 MO3aebfcgdh
+DNA鎖 , 2種, 2分子 NT
+Transcription initiation factor IIF subunit ... , 2種, 2分子 QR
+Transcription initiation factor IIA ... , 2種, 2分子 UV
+Transcription initiation factor IIE subunit ... , 2種, 2分子 WX
+General transcription and DNA repair factor IIH helicase subunit ... , 2種, 2分子 07
+General transcription and DNA repair factor IIH subunit ... , 5種, 5分子 12456
+非ポリマー , 3種, 19分子 




+詳細
-実験情報
-実験
| 実験 | 手法: 電子顕微鏡法 |
|---|---|
| EM実験 | 試料の集合状態: PARTICLE / 3次元再構成法: 単粒子再構成法 |
- 試料調製
試料調製
| 構成要素 | 名称: Yeast RNA polymerase II transcription pre-initiation complex with the +1 nucleosome タイプ: COMPLEX / Entity ID: #1-#28, #30-#34, #29 / 由来: MULTIPLE SOURCES |
|---|---|
| 分子量 | 値: 1.2 MDa / 実験値: NO |
| 由来(天然) | 生物種:  Saccharomyces killer virus M2-4 (ウイルス) Saccharomyces killer virus M2-4 (ウイルス) |
| 由来(組換発現) | 生物種:  |
| 緩衝液 | pH: 7.5 |
| 試料 | 包埋: NO / シャドウイング: NO / 染色: NO / 凍結: YES |
| 試料支持 | グリッドの材料: GOLD / グリッドのタイプ: UltrAuFoil R2/2 |
| 急速凍結 | 凍結剤: ETHANE |
- 電子顕微鏡撮影
電子顕微鏡撮影
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
|---|---|
| 顕微鏡 | モデル: FEI TITAN KRIOS |
| 電子銃 | 電子線源:  FIELD EMISSION GUN / 加速電圧: 300 kV / 照射モード: FLOOD BEAM FIELD EMISSION GUN / 加速電圧: 300 kV / 照射モード: FLOOD BEAM |
| 電子レンズ | モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2000 nm / 最小 デフォーカス(公称値): 800 nm |
| 撮影 | 電子線照射量: 42 e/Å2 フィルム・検出器のモデル: GATAN K3 BIOQUANTUM (6k x 4k) |
- 解析
解析
| ソフトウェア | 名称: PHENIX / バージョン: 1.19.2_4158: / 分類: 精密化 | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CTF補正 | タイプ: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||
| 3次元再構成 | 解像度: 3.1 Å / 解像度の算出法: FSC 0.143 CUT-OFF / 粒子像の数: 55851 / 対称性のタイプ: POINT | ||||||||||||||||||||||||
| 拘束条件 |
|
 ムービー
ムービー コントローラー
コントローラー



 PDBj
PDBj















































 Trichoplusia ni (イラクサキンウワバ)
Trichoplusia ni (イラクサキンウワバ)


