+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7zj4 | ||||||
|---|---|---|---|---|---|---|---|
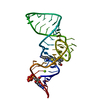
| Title | Ligand bound state of a brocolli-pepper aptamer FRET tile | ||||||
 Components Components | brocolli-pepper aptamer | ||||||
 Keywords Keywords | RNA / RNA origami aptamer fret | ||||||
| Function / homology | Chem-1TU / Chem-J93 / : / RNA / RNA (> 10) / RNA (> 100) Function and homology information Function and homology information | ||||||
| Biological species | synthetic construct (others) | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 4.43 Å | ||||||
 Authors Authors | McRae, E.K.S. / Vallina, N.S. / Hansen, B.K. / Boussebayle, A. / Andersen, E.S. | ||||||
| Funding support |  Denmark, 1items Denmark, 1items
| ||||||
 Citation Citation |  Journal: To Be Published Journal: To Be PublishedTitle: Structure determination of Pepper-Broccoli FRET pair by RNA origami scaffolding Authors: McRae, E.K.S. / Vallina, N.S. / Hansen, B.K. / Boussebayle, A. / Andersen, E.S. #1:  Journal: Acta Crystallogr D Struct Biol / Year: 2018 Journal: Acta Crystallogr D Struct Biol / Year: 2018Title: Rea-space refinement in PHENIX for cryo-EM and crystallography. Authors: Afonine, P.V. / Poon, B.K. / Read, R.J. / Sobolev, O.V. / Terwilliger, T.C. / Urzhumtsev, A. / Adams, P.D. #2:  Journal: Acta Crystallogr D Struct Biol / Year: 2018 Journal: Acta Crystallogr D Struct Biol / Year: 2018Title: ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Authors: Tristan Ian Croll /  Abstract: This paper introduces ISOLDE, a new software package designed to provide an intuitive environment for high-fidelity interactive remodelling/refinement of macromolecular models into electron-density ...This paper introduces ISOLDE, a new software package designed to provide an intuitive environment for high-fidelity interactive remodelling/refinement of macromolecular models into electron-density maps. ISOLDE combines interactive molecular-dynamics flexible fitting with modern molecular-graphics visualization and established structural biology libraries to provide an immersive interface wherein the model constantly acts to maintain physically realistic conformations as the user interacts with it by directly tugging atoms with a mouse or haptic interface or applying/removing restraints. In addition, common validation tasks are accelerated and visualized in real time. Using the recently described 3.8 Å resolution cryo-EM structure of the eukaryotic minichromosome maintenance (MCM) helicase complex as a case study, it is demonstrated how ISOLDE can be used alongside other modern refinement tools to avoid common pitfalls of low-resolution modelling and improve the quality of the final model. A detailed analysis of changes between the initial and final model provides a somewhat sobering insight into the dangers of relying on a small number of validation metrics to judge the quality of a low-resolution model. #3:  Journal: Nat Methods / Year: 2017 Journal: Nat Methods / Year: 2017Title: cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Authors: Punjani, A. / Rubinstein, J.L. / Fleet, D.J. / Brubaker, M.A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7zj4.cif.gz 7zj4.cif.gz | 273.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7zj4.ent.gz pdb7zj4.ent.gz | 204.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7zj4.json.gz 7zj4.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  7zj4_validation.pdf.gz 7zj4_validation.pdf.gz | 1.2 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  7zj4_full_validation.pdf.gz 7zj4_full_validation.pdf.gz | 1.2 MB | Display | |
| Data in XML |  7zj4_validation.xml.gz 7zj4_validation.xml.gz | 23.8 KB | Display | |
| Data in CIF |  7zj4_validation.cif.gz 7zj4_validation.cif.gz | 34.1 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/zj/7zj4 https://data.pdbj.org/pub/pdb/validation_reports/zj/7zj4 ftp://data.pdbj.org/pub/pdb/validation_reports/zj/7zj4 ftp://data.pdbj.org/pub/pdb/validation_reports/zj/7zj4 | HTTPS FTP |
-Related structure data
| Related structure data |  14740MC  7zj5C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: RNA chain | Mass: 120700.953 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) |
|---|---|
| #2: Chemical | ChemComp-J93 / |
| #3: Chemical | ChemComp-1TU / |
| #4: Chemical | ChemComp-K / |
| Has ligand of interest | Y |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Ligand bound state of a brocolli-pepper aptamer FRET tile Type: COMPLEX / Entity ID: #1 / Source: RECOMBINANT |
|---|---|
| Molecular weight | Value: 0.12 MDa / Experimental value: NO |
| Source (natural) | Organism: synthetic construct (others) |
| Source (recombinant) | Organism: synthetic construct (others) |
| Buffer solution | pH: 7.5 Details: 40mM HEPES pH 7.5, 5mM MgCl2, 50mM KCl. Filtered through 0.22 um filter. |
| Specimen | Conc.: 2.5 mg/ml / Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES Details: Sample was purified by size exclusion chromatography. |
| Specimen support | Details: 15mA of current. / Grid material: GOLD / Grid mesh size: 300 divisions/in. / Grid type: UltrAuFoil R1.2/1.3 |
| Vitrification | Instrument: LEICA EM GP / Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 288 K Details: 3 uL sample, blotted onto double layer of whatman filter paper for 6 seconds. |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal defocus max: 2000 nm / Nominal defocus min: 700 nm / Cs: 2.7 mm |
| Image recording | Electron dose: 60 e/Å2 / Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Num. of grids imaged: 1 / Num. of real images: 5354 Details: Collected with a calibrated pixel size of 0.647 Angstrom |
| EM imaging optics | Energyfilter name: GIF Bioquantum / Energyfilter slit width: 20 eV |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||||||||||||||||||||||||||
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 729630 Details: Picked from templates generated from an ab initio model | ||||||||||||||||||||||||||||||||||||
| Symmetry | Point symmetry: C1 (asymmetric) | ||||||||||||||||||||||||||||||||||||
| 3D reconstruction | Resolution: 4.43 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 150204 Details: Local refinement was performed using a mask covering the entire volume. Num. of class averages: 1 / Symmetry type: POINT | ||||||||||||||||||||||||||||||||||||
| Atomic model building | Protocol: FLEXIBLE FIT / Space: REAL | ||||||||||||||||||||||||||||||||||||
| Atomic model building |
| ||||||||||||||||||||||||||||||||||||
| Refinement | Cross valid method: NONE | ||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller




 PDBj
PDBj