+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9852 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of a Q-engaged arrested complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | RNA polymerase / Antitermination / TRANSCRIPTION / DNA / RNA | |||||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of termination of DNA-templated transcription / sigma factor antagonist complex / RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / regulation of DNA-templated transcription initiation / sigma factor activity / bacterial-type flagellum assembly / bacterial-type RNA polymerase core enzyme binding / cytosolic DNA-directed RNA polymerase complex ...negative regulation of termination of DNA-templated transcription / sigma factor antagonist complex / RNA polymerase complex / submerged biofilm formation / cellular response to cell envelope stress / regulation of DNA-templated transcription initiation / sigma factor activity / bacterial-type flagellum assembly / bacterial-type RNA polymerase core enzyme binding / cytosolic DNA-directed RNA polymerase complex / bacterial-type flagellum-dependent cell motility / nitrate assimilation / regulation of DNA-templated transcription elongation / transcription elongation factor complex / transcription antitermination / cell motility / DNA-templated transcription initiation / ribonucleoside binding / DNA-directed RNA polymerase / DNA-directed RNA polymerase activity / response to heat / protein-containing complex assembly / intracellular iron ion homeostasis / protein dimerization activity / response to antibiotic / negative regulation of DNA-templated transcription / magnesium ion binding / DNA binding / zinc ion binding / membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   Enterobacteria phage SfI (virus) / Enterobacteria phage SfI (virus) /  Enterobacteria phage P21 (virus) Enterobacteria phage P21 (virus) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.08 Å | |||||||||
 Authors Authors | Feng Y / Shi J | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2019 Journal: Nat Commun / Year: 2019Title: Structural basis of Q-dependent transcription antitermination. Authors: Jing Shi / Xiang Gao / Tongguan Tian / Zhaoyang Yu / Bo Gao / Aijia Wen / Linlin You / Shenghai Chang / Xing Zhang / Yu Zhang / Yu Feng /  Abstract: Bacteriophage Q protein engages σ-dependent paused RNA polymerase (RNAP) by binding to a DNA site embedded in late gene promoter and renders RNAP resistant to termination signals. Here, we report a ...Bacteriophage Q protein engages σ-dependent paused RNA polymerase (RNAP) by binding to a DNA site embedded in late gene promoter and renders RNAP resistant to termination signals. Here, we report a single-particle cryo-electron microscopy (cryo-EM) structure of an intact Q-engaged arrested complex. The structure reveals key interactions responsible for σ-dependent pause, Q engagement, and Q-mediated transcription antitermination. The structure shows that two Q protomers (Q and Q) bind to a direct-repeat DNA site and contact distinct elements of the RNA exit channel. Notably, Q forms a narrow ring inside the RNA exit channel and renders RNAP resistant to termination signals by prohibiting RNA hairpin formation in the RNA exit channel. Because the RNA exit channel is conserved among all multisubunit RNAPs, it is likely to serve as an important contact site for regulators that modify the elongation properties of RNAP in other organisms, as well. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9852.map.gz emd_9852.map.gz | 28.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9852-v30.xml emd-9852-v30.xml emd-9852.xml emd-9852.xml | 21.7 KB 21.7 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9852.png emd_9852.png | 65.4 KB | ||
| Filedesc metadata |  emd-9852.cif.gz emd-9852.cif.gz | 8.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9852 http://ftp.pdbj.org/pub/emdb/structures/EMD-9852 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9852 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9852 | HTTPS FTP |
-Validation report
| Summary document |  emd_9852_validation.pdf.gz emd_9852_validation.pdf.gz | 597.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_9852_full_validation.pdf.gz emd_9852_full_validation.pdf.gz | 597.5 KB | Display | |
| Data in XML |  emd_9852_validation.xml.gz emd_9852_validation.xml.gz | 5.6 KB | Display | |
| Data in CIF |  emd_9852_validation.cif.gz emd_9852_validation.cif.gz | 6.4 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9852 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9852 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9852 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9852 | HTTPS FTP |
-Related structure data
| Related structure data |  6jnxMC  6jnyC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9852.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9852.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.307 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
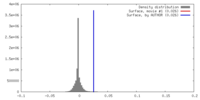
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : 21Q-engaged arrested complex
+Supramolecule #1: 21Q-engaged arrested complex
+Macromolecule #1: DNA-directed RNA polymerase subunit alpha
+Macromolecule #2: DNA-directed RNA polymerase subunit beta
+Macromolecule #3: DNA-directed RNA polymerase subunit beta'
+Macromolecule #4: DNA-directed RNA polymerase subunit omega
+Macromolecule #5: RNA polymerase sigma factor RpoD
+Macromolecule #9: Antiterminator Q protein
+Macromolecule #6: DNA (63-MER)
+Macromolecule #8: DNA (63-MER)
+Macromolecule #7: RNA (5'-R(P*AP*UP*AP*AP*GP*GP*UP*GP*GP*GP*GP*UP*UP*AP*GP*UP*GP*A)-3')
+Macromolecule #10: MAGNESIUM ION
+Macromolecule #11: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 8 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 283 K |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 56.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: EMDB MAP |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 4.08 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 64497 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
 Movie
Movie Controller
Controller
















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)