+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8579 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | membrane protein | |||||||||
 Map data Map data | original map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | membrane protein | |||||||||
| Function / homology |  Function and homology information Function and homology informationNMDA glutamate receptor activity / NMDA selective glutamate receptor complex / ligand-gated sodium channel activity / response to zinc ion / calcium ion transmembrane import into cytosol / protein heterotetramerization / response to magnesium ion / glutamate-gated calcium ion channel activity / ionotropic glutamate receptor signaling pathway / sodium ion transmembrane transport ...NMDA glutamate receptor activity / NMDA selective glutamate receptor complex / ligand-gated sodium channel activity / response to zinc ion / calcium ion transmembrane import into cytosol / protein heterotetramerization / response to magnesium ion / glutamate-gated calcium ion channel activity / ionotropic glutamate receptor signaling pathway / sodium ion transmembrane transport / regulation of membrane potential / excitatory postsynaptic potential / synaptic transmission, glutamatergic / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / regulation of synaptic plasticity / postsynaptic density membrane / calcium ion transmembrane transport / long-term synaptic potentiation / late endosome / signaling receptor activity / chemical synaptic transmission / postsynaptic membrane / lysosome / neuron projection / synapse / metal ion binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.5 Å | |||||||||
 Authors Authors | Lu W / Du J / Goehring A / Gouaux E | |||||||||
 Citation Citation |  Journal: Science / Year: 2017 Journal: Science / Year: 2017Title: Cryo-EM structures of the triheteromeric NMDA receptor and its allosteric modulation. Authors: Wei Lü / Juan Du / April Goehring / Eric Gouaux /  Abstract: -methyl-d-aspartate receptors (NMDARs) are heterotetrameric ion channels assembled as diheteromeric or triheteromeric complexes. Here, we report structures of the triheteromeric GluN1/GluN2A/GluN2B ...-methyl-d-aspartate receptors (NMDARs) are heterotetrameric ion channels assembled as diheteromeric or triheteromeric complexes. Here, we report structures of the triheteromeric GluN1/GluN2A/GluN2B receptor in the absence or presence of the GluN2B-specific allosteric modulator Ro 25-6981 (Ro), determined by cryogenic electron microscopy (cryo-EM). In the absence of Ro, the GluN2A and GluN2B amino-terminal domains (ATDs) adopt "closed" and "open" clefts, respectively. Upon binding Ro, the GluN2B ATD clamshell transitions from an open to a closed conformation. Consistent with a predominance of the GluN2A subunit in ion channel gating, the GluN2A subunit interacts more extensively with GluN1 subunits throughout the receptor, in comparison with the GluN2B subunit. Differences in the conformation of the pseudo-2-fold-related GluN1 subunits further reflect receptor asymmetry. The triheteromeric NMDAR structures provide the first view of the most common NMDA receptor assembly and show how incorporation of two different GluN2 subunits modifies receptor symmetry and subunit interactions, allowing each subunit to uniquely influence receptor structure and function, thus increasing receptor complexity. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8579.map.gz emd_8579.map.gz | 10.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8579-v30.xml emd-8579-v30.xml emd-8579.xml emd-8579.xml | 18 KB 18 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_8579.png emd_8579.png | 166.7 KB | ||
| Filedesc metadata |  emd-8579.cif.gz emd-8579.cif.gz | 6.7 KB | ||
| Others |  emd_8579_additional.map.gz emd_8579_additional.map.gz | 4.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8579 http://ftp.pdbj.org/pub/emdb/structures/EMD-8579 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8579 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8579 | HTTPS FTP |
-Related structure data
| Related structure data |  5uowMC  8580C  8581C  8583C  5up2C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8579.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8579.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
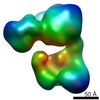
| Annotation | original map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.7 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: sharpened map
| File | emd_8579_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : membrane protein
| Entire | Name: membrane protein |
|---|---|
| Components |
|
-Supramolecule #1: membrane protein
| Supramolecule | Name: membrane protein / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#4 |
|---|---|
| Source (natural) | Organism: |
-Macromolecule #1: N-methyl-D-aspartate receptor subunit NR1-8a
| Macromolecule | Name: N-methyl-D-aspartate receptor subunit NR1-8a / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 91.639867 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DPKIVNIGAV LSTKKHEQIF REAVNQANKR HFTRKIQLNA TSVTHRPNAI QMALSVCEDL ISSQVYAILV SHPPAPTDHL TPTPISYTA GFYRIPVIGL TTRMSIYSDK SIHLSFLRTV PPYSHQALVW FEMMRLFNWN HVILIVSDDH EGRAAQKKLE T LLEEKESK ...String: DPKIVNIGAV LSTKKHEQIF REAVNQANKR HFTRKIQLNA TSVTHRPNAI QMALSVCEDL ISSQVYAILV SHPPAPTDHL TPTPISYTA GFYRIPVIGL TTRMSIYSDK SIHLSFLRTV PPYSHQALVW FEMMRLFNWN HVILIVSDDH EGRAAQKKLE T LLEEKESK ADKVLQFEPG TKNLTALLLE AKELEARVII LSASEDDATA VYKSAAMLDM TGAGYVWLVG EREISGSALR YA PDGIIGL QLINGKNESA HISDAVAVVA QAIHELFEME QITDPPRGCV GNTNIWKTGP LFKRVLMSSK YPDGVTGRIE FNE DGDRKF AQYSIMNLQN RKLVQVGIFD GSYIIQNDRK IIWPGGETER PQGYQMSTRL KIVTIHQEPF VYVRPTTSDG TCRE EYTIN GDPIKKVICN GPDETIPGRP TVPQCCYGFC VDLLIKLARE MDFTYEVHLV ADGKFGTQER VNNSNAAAWN GMMGE LLSG QADMIVAPLT INNERAQYIE FSKPFKYQGL TILVKKEIPR STLDSFMQPF QSTLWLLVGL SVHVVAVMLY LLDRFS PFG RFKVNSAAAE EDALTLSSAM WFSWRVLLNS GLGEGAPRSF SARILGMVWA LFAMIIVASY TANLAAFLVL RRPEERI TG INDPRLRNPS DKFIYATVKQ SSVDIYFRRQ VELSTMYRHM EKHNYESAAE AIQAVRDNKL HAFIWDSAVL EFEASQDC D LVTTGELFFR SGFGIGMRKD SPWKQEVSLN ILKSHENGFM EELDKTWVRY QECDSRSNAP ATLTFENMAG VFYLVAGGI VAGIFLIFIE IAYK UniProtKB: Glutamate receptor ionotropic, NMDA 1 |
-Macromolecule #2: N-methyl-D-aspartate receptor subunit NR2A
| Macromolecule | Name: N-methyl-D-aspartate receptor subunit NR2A / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 93.664344 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGMFVLLLYT FLYAGDLGHG AEKSFPVLNI AVILGRTRYI TERDIRSLWT RDMSLDFDVN VVTLLVQQTD PKSIITHVCD LMSGTKIHG VVFGDDTDQE AIAQILDFVS SQTFIPILGI HGGSSMIMAD KDEMSTFFQF GASIKQQATV MLNIMEEYDW H VFSVITSN ...String: MGMFVLLLYT FLYAGDLGHG AEKSFPVLNI AVILGRTRYI TERDIRSLWT RDMSLDFDVN VVTLLVQQTD PKSIITHVCD LMSGTKIHG VVFGDDTDQE AIAQILDFVS SQTFIPILGI HGGSSMIMAD KDEMSTFFQF GASIKQQATV MLNIMEEYDW H VFSVITSN FPGYRDFISF IKTTVDNSFV GWEVQNYITL DTSYTDAQTL TQLKKIHSSV ILLYCSKDEA TYIFEEARSL GL MGYGFVW IVPSLVTGNT DIIPYEFPSG LVSVSYDDWD YGIEARVRDG LGIITTAASA MLEKHSVIPE AKTSCYGQNE RND PPLHTL HNFMINVTWD GKDLSFTEDG YQANPKLVVL LLNMEREWEK VGKWEAKSLN MKYPVWPRID SDHDDNHLSI VTLE EAPFV IVENIDYLTG TCVRNTVPCR KYFRLANSTT EGTSVKKCCK GFCIDILKKL SKTVKFTYDL YLVTNGKHGK KIKNV WNGM IGEVVYKRAV MAVGSLTINE ERSVAVDFSV PFVETGISVM VSRSAGTVSP SAFLEPFSAS VWVMMFVMLL LVSAMA VFI FEYFSPVGYN RNLAQGKDPH GPSFTIGKAV WLLWGLVFNN SLPVQNPKGT TSKIIVSIWA FFAVIFLASY TANLAAF MI QRRFVDQVTG LSDNKFQRPH DYSPPFRFGT VPQGSTERNI RNNYPDMHQY MVKFHQKGVQ DALVSLKTGK LDAFIYDA A VLNYMAGRDE GCKLVTIGSG YIFATTGYGI ALQKGSRWKR PIDLALLQFV GDGEMEELEK LWLTGICHTE KNEVMSSQL DIDNMAGVFY MLAAAMALSL ITFVWEHLFY KE UniProtKB: Glutamate receptor ionotropic, NMDA 2A |
-Macromolecule #3: Ionotropic glutamate receptor subunit NR2B
| Macromolecule | Name: Ionotropic glutamate receptor subunit NR2B / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 94.552234 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MRPTEACCYL KISLIILFYS RAYAQKHPNM DIAVILVGTT EEVAIKDVHE KDDFHHLPVT PRVELVTMQE SDPKSIITRI CDLMSDKKV QGVVFGDDTD QEAIAQILDF ISVQTLTPIL GIHGGSSMIM ADKEEASMFF QFGPSIEQQA SVMLNIMEEY D WYIFSIVT ...String: MRPTEACCYL KISLIILFYS RAYAQKHPNM DIAVILVGTT EEVAIKDVHE KDDFHHLPVT PRVELVTMQE SDPKSIITRI CDLMSDKKV QGVVFGDDTD QEAIAQILDF ISVQTLTPIL GIHGGSSMIM ADKEEASMFF QFGPSIEQQA SVMLNIMEEY D WYIFSIVT TYFPGYQDFE NKVRSTIENS FVGWELEEVI HLDMSLDDID SKIQNQLKKL QSPVILLYCT KEEATYIFEV AH SVGLTGY GFTWIVPSLV AGDTDTVPDE FPTGLISVSY DEWDYDLPAR VRDGIAIITT AASTMLSEHN SIPQSKSSCN NIQ ESRVYE AHMLKRYLIN VTFEGRDLSF SEDGYQMHPK LVIILLNQER KWERVGKYKD RSLKMWPVFD LYPNSEEHKD EHLS IVTLE EAPFVIVEDV DPLSGTCMRN TVPCRKQIRP ENRTEEGGNY IKRCCKGFCI DILKKIAKTV KFTYDLYLVT NGKHG KKIN GVWNGMIGEV VTKRAYMAVG SLTINEERSE VVDFSVPFIE TGISVMVSRS NGTVSPSAFL EPFSADVWVM MFVMLL IVS AVAVFVFEYF SPVGYNRALA DGREPGGPSF TIGKAIWLLW GLVFNNSLPV QNPKGTTSKI MVSVWAFFAV IFLASYT AN LAAFMIQRRY VDQVSGLSDK KFQRPNDFSP AFRFGTVPNG STERNIRNNY LEMHSYMVKF NQRSVQDALL SLKSGKLD A FIYDAAVLNY MAGRDEGCKL VTIGSGKVFA TTGYGIAIQK DSGWKRQVDL AILQLFGDGE MEELEALWLT GICHNEKNE VMSSQLDIDN MAGVFYMLAA AMALSLITFI MEHLFYK UniProtKB: Glutamate receptor ionotropic, NMDA 2B |
-Macromolecule #4: GluN2B-specific Fab, termed 11D1
| Macromolecule | Name: GluN2B-specific Fab, termed 11D1 / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 18.400619 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) |
-Macromolecule #6: GLUTAMIC ACID
| Macromolecule | Name: GLUTAMIC ACID / type: ligand / ID: 6 / Number of copies: 1 / Formula: GLU |
|---|---|
| Molecular weight | Theoretical: 147.129 Da |
| Chemical component information |  ChemComp-GLU: |
-Macromolecule #7: (5S,10R)-5-methyl-10,11-dihydro-5H-5,10-epiminodibenzo[a,d][7]annulene
| Macromolecule | Name: (5S,10R)-5-methyl-10,11-dihydro-5H-5,10-epiminodibenzo[a,d][7]annulene type: ligand / ID: 7 / Number of copies: 1 / Formula: BMK |
|---|---|
| Molecular weight | Theoretical: 221.297 Da |
| Chemical component information | 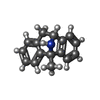 ChemComp-BMK: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 6.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 0.84 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: PDB ENTRY |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 4.5 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 302052 |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION |
| Final angle assignment | Type: ANGULAR RECONSTITUTION |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)