[English] 日本語
 Yorodumi
Yorodumi- EMDB-8301: Architecture of the Human Mitochondrial Iron-Sulfur Cluster Assem... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8301 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Architecture of the Human Mitochondrial Iron-Sulfur Cluster Assembly Machinery: the Complex Formed by the Iron Donor, the Sulfur Donor, and the Scaffold | |||||||||
 Map data Map data | Human mitochondrial iron-sulfur cluster assembly machinery: grouping of segments corresponding to one half of the full map (EMD-8293) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | frataxin / iron-sulfur protein / mitochondria / protein complex / TRANSFERASE-OXIDOREDUCTASE complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of lyase activity / iron-sulfur cluster chaperone activity / negative regulation of iron ion import across plasma membrane / molybdopterin cofactor metabolic process / Molybdenum cofactor biosynthesis / proprioception / [4Fe-4S] cluster assembly / Mitochondrial iron-sulfur cluster biogenesis / sulfur carrier activity / Complex III assembly ...positive regulation of lyase activity / iron-sulfur cluster chaperone activity / negative regulation of iron ion import across plasma membrane / molybdopterin cofactor metabolic process / Molybdenum cofactor biosynthesis / proprioception / [4Fe-4S] cluster assembly / Mitochondrial iron-sulfur cluster biogenesis / sulfur carrier activity / Complex III assembly / iron chaperone activity / positive regulation of mitochondrial electron transport, NADH to ubiquinone / Maturation of TCA enzymes and regulation of TCA cycle / cysteine desulfurase / negative regulation of organ growth / cysteine desulfurase activity / mitochondrial respiratory chain complex III assembly / Mo-molybdopterin cofactor biosynthetic process / embryo development ending in birth or egg hatching / Mitochondrial protein import / iron-sulfur cluster assembly complex / mitochondrial [2Fe-2S] assembly complex / response to iron ion / oxidative phosphorylation / adult walking behavior / heme biosynthetic process / [2Fe-2S] cluster assembly / negative regulation of multicellular organism growth / organ growth / muscle cell cellular homeostasis / iron-sulfur cluster assembly / negative regulation of release of cytochrome c from mitochondria / ferroxidase / protein autoprocessing / ferroxidase activity / iron-sulfur cluster binding / ferric iron binding / protein maturation / iron ion transport / enzyme activator activity / ferrous iron binding / 2 iron, 2 sulfur cluster binding / cellular response to hydrogen peroxide / pyridoxal phosphate binding / Maturation of replicase proteins / molecular adaptor activity / intracellular iron ion homeostasis / iron ion binding / mitochondrial matrix / centrosome / negative regulation of apoptotic process / protein homodimerization activity / mitochondrion / zinc ion binding / nucleoplasm / metal ion binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
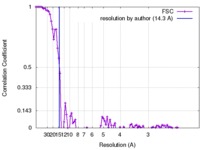
| Method | single particle reconstruction / negative staining / Resolution: 14.3 Å | |||||||||
 Authors Authors | Gakh O / Ranatunga W / Smith DY / Ahlgren EC / Al-Karadaghi S / Thompson JR / Isaya G | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: J Biol Chem / Year: 2016 Journal: J Biol Chem / Year: 2016Title: Architecture of the Human Mitochondrial Iron-Sulfur Cluster Assembly Machinery. Authors: Oleksandr Gakh / Wasantha Ranatunga / Douglas Y Smith / Eva-Christina Ahlgren / Salam Al-Karadaghi / James R Thompson / Grazia Isaya /   Abstract: Fe-S clusters, essential cofactors needed for the activity of many different enzymes, are assembled by conserved protein machineries inside bacteria and mitochondria. As the architecture of the human ...Fe-S clusters, essential cofactors needed for the activity of many different enzymes, are assembled by conserved protein machineries inside bacteria and mitochondria. As the architecture of the human machinery remains undefined, we co-expressed in Escherichia coli the following four proteins involved in the initial step of Fe-S cluster synthesis: FXN (iron donor); [NFS1]·[ISD11] (sulfur donor); and ISCU (scaffold upon which new clusters are assembled). We purified a stable, active complex consisting of all four proteins with 1:1:1:1 stoichiometry. Using negative staining transmission EM and single particle analysis, we obtained a three-dimensional model of the complex with ∼14 Å resolution. Molecular dynamics flexible fitting of protein structures docked into the EM map of the model revealed a [FXN]·[NFS1]·[ISD11]·[ISCU] complex, consistent with the measured 1:1:1:1 stoichiometry of its four components. The complex structure fulfills distance constraints obtained from chemical cross-linking of the complex at multiple recurring interfaces, involving hydrogen bonds, salt bridges, or hydrophobic interactions between conserved residues. The complex consists of a central roughly cubic [FXN]·[ISCU] sub-complex with one symmetric ISCU trimer bound on top of one symmetric FXN trimer at each of its eight vertices. Binding of 12 [NFS1]·[ISD11] sub-complexes to the surface results in a globular macromolecule with a diameter of ∼15 nm and creates 24 Fe-S cluster assembly centers. The organization of each center recapitulates a previously proposed conserved mechanism for sulfur donation from NFS1 to ISCU and reveals, for the first time, a path for iron donation from FXN to ISCU. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8301.map.gz emd_8301.map.gz | 1.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8301-v30.xml emd-8301-v30.xml emd-8301.xml emd-8301.xml | 15.5 KB 15.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8301_fsc.xml emd_8301_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_8301.png emd_8301.png | 85.3 KB | ||
| Filedesc metadata |  emd-8301.cif.gz emd-8301.cif.gz | 6.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8301 http://ftp.pdbj.org/pub/emdb/structures/EMD-8301 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8301 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8301 | HTTPS FTP |
-Related structure data
| Related structure data |  5kz5MC  8293C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8301.map.gz / Format: CCP4 / Size: 11.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8301.map.gz / Format: CCP4 / Size: 11.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Human mitochondrial iron-sulfur cluster assembly machinery: grouping of segments corresponding to one half of the full map (EMD-8293) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : NFS1-ISD11-ISCU-FXN
| Entire | Name: NFS1-ISD11-ISCU-FXN |
|---|---|
| Components |
|
-Supramolecule #1: NFS1-ISD11-ISCU-FXN
| Supramolecule | Name: NFS1-ISD11-ISCU-FXN / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Cysteine desulfurase, mitochondrial
| Macromolecule | Name: Cysteine desulfurase, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 12 / Enantiomer: LEVO / EC number: cysteine desulfurase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 43.429648 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: TPLDPRVLDA MLPYLINYYG NPHSRTHAYG WESEAAMERA RQQVASLIGA DPREIIFTSG ATESNNIAIK GVARFYRSRK KHLITTQTE HKCVLDSCRS LEAEGFQVTY LPVQKSGIID LKELEAAIQP DTSLVSVMTV NNEIGVKQPI AEIGRICSSR K VYFHTDAA ...String: TPLDPRVLDA MLPYLINYYG NPHSRTHAYG WESEAAMERA RQQVASLIGA DPREIIFTSG ATESNNIAIK GVARFYRSRK KHLITTQTE HKCVLDSCRS LEAEGFQVTY LPVQKSGIID LKELEAAIQP DTSLVSVMTV NNEIGVKQPI AEIGRICSSR K VYFHTDAA QAVGKIPLDV NDMKIDLMSI SGHKIYGPKG VGAIYIRRRP RVRVEALQSG GGQERGMRSG TVPTPLVVGL GA ACEVAQQ EMEYDHKRIS KLSERLIQNI MKSLPDVVMN GDPKHHYPGC INLSFAYVEG ESLLMALKDV ALSSGSACTS ASL EPSYVL RAIGTDEDLA HSSIRFGIGR FTTEEEVDYT VEKCIQHVKR LREMSPLWEM VQDGIDLKSI KWTQH UniProtKB: Cysteine desulfurase |
-Macromolecule #2: Frataxin, mitochondrial
| Macromolecule | Name: Frataxin, mitochondrial / type: protein_or_peptide / ID: 2 / Number of copies: 12 / Enantiomer: LEVO / EC number: ferroxidase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 18.849025 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: LRTDIDATCT PRRASSNQRG LNQIWNVKKQ SVYLMNLRKS GTLGHPGSLD ETTYERLAEE TLDSLAEFFE DLADKPYTFE DYDVSFGSG VLTVKLGGDL GTYVINKQTP NKQIWLSSPS SGPKRYDWTG KNWVYSHDGV SLHELLAAEL TKALKTKLDL S SLAYSGKD A UniProtKB: Frataxin, mitochondrial |
-Macromolecule #3: Iron-sulfur cluster assembly enzyme ISCU, mitochondrial
| Macromolecule | Name: Iron-sulfur cluster assembly enzyme ISCU, mitochondrial type: protein_or_peptide / ID: 3 / Number of copies: 12 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 12.52558 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GSLDKTSKNV GTGLVGAPAC GDVMKLQIQV DEKGKIVDAR FKTFGCGSAI ASSSLATEWV KGKTVEEALT IKNTDIAKEL CLPPVKLHC SMLAEDAIKA ALADYKLKQE PKKGEAEKK UniProtKB: Iron-sulfur cluster assembly enzyme ISCU |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| |||||||||
| Staining | Type: NEGATIVE / Material: 1% uranyl acetate / Details: 5 and 30 seconds | |||||||||
| Grid | Model: Carbon-coated copper grids, EMS / Material: COPPER / Mesh: 400 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Details: DV-502A vacuum evaporator (Denton Vacuum Inc.) | |||||||||
| Details | 50 mM Tris-HCl, pH 8.0, 150 mM NaCl |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F30 |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 466 / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 3.0 µm / Calibrated defocus min: 0.21 µm / Calibrated magnification: 115000 / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.0 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 0.21 µm / Nominal magnification: 115000 |
| Sample stage | Specimen holder model: SIDE ENTRY, EUCENTRIC / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
|---|---|
| Output model |  PDB-5kz5: |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)