[English] 日本語
 Yorodumi
Yorodumi- EMDB-7821: CryoEM structure of Tetrahymena telomerase with telomeric DNA at ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7821 | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of Tetrahymena telomerase with telomeric DNA at 4.8 Angstrom resolution | ||||||||||||||||||||||||||||||
 Map data Map data | Tetrahymena telomerase with telomeric DNA | ||||||||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||||||||
 Keywords Keywords | Telomerase / telomere / REPLICATION | ||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationtelomerase catalytic core complex assembly / telomerase RNA stabilization / telomerase catalytic core complex / DNA replication factor A complex / telomerase activity / single-stranded telomeric DNA binding / telomerase holoenzyme complex / telomerase RNA binding / telomeric DNA binding / telomere maintenance via telomerase ...telomerase catalytic core complex assembly / telomerase RNA stabilization / telomerase catalytic core complex / DNA replication factor A complex / telomerase activity / single-stranded telomeric DNA binding / telomerase holoenzyme complex / telomerase RNA binding / telomeric DNA binding / telomere maintenance via telomerase / RNA-directed DNA polymerase / DNA recombination / DNA replication / chromosome, telomeric region / DNA repair / DNA binding / zinc ion binding / metal ion binding Similarity search - Function | ||||||||||||||||||||||||||||||
| Biological species |  | ||||||||||||||||||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.8 Å | ||||||||||||||||||||||||||||||
 Authors Authors | Jiang J / Wang Y / Susac L / Chan H / Basu R / Zhou ZH / Feigon J | ||||||||||||||||||||||||||||||
| Funding support |  United States, 9 items United States, 9 items
| ||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Cell / Year: 2018 Journal: Cell / Year: 2018Title: Structure of Telomerase with Telomeric DNA. Authors: Jiansen Jiang / Yaqiang Wang / Lukas Sušac / Henry Chan / Ritwika Basu / Z Hong Zhou / Juli Feigon /  Abstract: Telomerase is an RNA-protein complex (RNP) that extends telomeric DNA at the 3' ends of chromosomes using its telomerase reverse transcriptase (TERT) and integral template-containing telomerase RNA ...Telomerase is an RNA-protein complex (RNP) that extends telomeric DNA at the 3' ends of chromosomes using its telomerase reverse transcriptase (TERT) and integral template-containing telomerase RNA (TER). Its activity is a critical determinant of human health, affecting aging, cancer, and stem cell renewal. Lack of atomic models of telomerase, particularly one with DNA bound, has limited our mechanistic understanding of telomeric DNA repeat synthesis. We report the 4.8 Å resolution cryoelectron microscopy structure of active Tetrahymena telomerase bound to telomeric DNA. The catalytic core is an intricately interlocked structure of TERT and TER, including a previously structurally uncharacterized TERT domain that interacts with the TEN domain to physically enclose TER and regulate activity. This complete structure of a telomerase catalytic core and its interactions with telomeric DNA from the template to telomere-interacting p50-TEB complex provides unanticipated insights into telomerase assembly and catalytic cycle and a new paradigm for a reverse transcriptase RNP. | ||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7821.map.gz emd_7821.map.gz | 58.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7821-v30.xml emd-7821-v30.xml emd-7821.xml emd-7821.xml | 35.5 KB 35.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_7821.png emd_7821.png | 153.8 KB | ||
| Masks |  emd_7821_msk_1.map emd_7821_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-7821.cif.gz emd-7821.cif.gz | 9.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7821 http://ftp.pdbj.org/pub/emdb/structures/EMD-7821 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7821 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7821 | HTTPS FTP |
-Related structure data
| Related structure data |  6d6vMC  7820C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_7821.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7821.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Tetrahymena telomerase with telomeric DNA | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.36 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
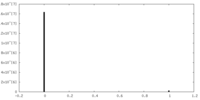
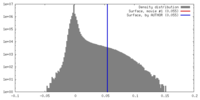
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_7821_msk_1.map emd_7821_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
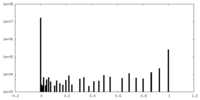
| Density Histograms |
- Sample components
Sample components
+Entire : Endogenous assembled Tetrahymena telomerase bound with telomeric DNA
+Supramolecule #1: Endogenous assembled Tetrahymena telomerase bound with telomeric DNA
+Macromolecule #1: Telomerase-associated protein 82
+Macromolecule #2: Telomerase reverse transcriptase
+Macromolecule #3: Telomerase holoenzyme TEB heterotrimer Teb3 subunit
+Macromolecule #4: Telomerase holoenzyme Teb2 subunit
+Macromolecule #6: Telomerase-associated protein 50
+Macromolecule #7: Telomerase La-related protein p65
+Macromolecule #5: RNA (159-MER)
+Macromolecule #8: DNA (5'-D(P*GP*TP*TP*GP*GP*GP*GP*TP*TP*GP*GP*GP*GP*TP*TP*GP*GP*GP...
+Macromolecule #9: ZINC ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average exposure time: 12.0 sec. / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)