+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6911 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of human TRPC3 at 4.36A resolution | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | TRPC3 channel / MEMBRANE PROTEIN | ||||||||||||
| Function / homology | :  Function and homology information Function and homology information | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.36 Å | ||||||||||||
 Authors Authors | Chen L / Tang Q | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell Res / Year: 2018 Journal: Cell Res / Year: 2018Title: Structure of the receptor-activated human TRPC6 and TRPC3 ion channels. Authors: Qinglin Tang / Wenjun Guo / Li Zheng / Jing-Xiang Wu / Meng Liu / Xindi Zhou / Xiaolin Zhang / Lei Chen /  Abstract: TRPC6 and TRPC3 are receptor-activated nonselective cation channels that belong to the family of canonical transient receptor potential (TRPC) channels. They are activated by diacylglycerol, a lipid ...TRPC6 and TRPC3 are receptor-activated nonselective cation channels that belong to the family of canonical transient receptor potential (TRPC) channels. They are activated by diacylglycerol, a lipid second messenger. TRPC6 and TRPC3 are involved in many physiological processes and implicated in human genetic diseases. Here we present the structure of human TRPC6 homotetramer in complex with a newly identified high-affinity inhibitor BTDM solved by single-particle cryo-electron microscopy to 3.8 Å resolution. We also present the structure of human TRPC3 at 4.4 Å resolution. These structures show two-layer architectures in which the bell-shaped cytosolic layer holds the transmembrane layer. Extensive inter-subunit interactions of cytosolic domains, including the N-terminal ankyrin repeats and the C-terminal coiled-coil, contribute to the tetramer assembly. The high-affinity inhibitor BTDM wedges between the S5-S6 pore domain and voltage sensor-like domain to inhibit channel opening. Our structures uncover the molecular architecture of TRPC channels and provide a structural basis for understanding the mechanism of these channels. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6911.map.gz emd_6911.map.gz | 28.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6911-v30.xml emd-6911-v30.xml emd-6911.xml emd-6911.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6911.png emd_6911.png | 133.2 KB | ||
| Filedesc metadata |  emd-6911.cif.gz emd-6911.cif.gz | 5.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6911 http://ftp.pdbj.org/pub/emdb/structures/EMD-6911 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6911 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6911 | HTTPS FTP |
-Validation report
| Summary document |  emd_6911_validation.pdf.gz emd_6911_validation.pdf.gz | 541.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6911_full_validation.pdf.gz emd_6911_full_validation.pdf.gz | 541.1 KB | Display | |
| Data in XML |  emd_6911_validation.xml.gz emd_6911_validation.xml.gz | 5.8 KB | Display | |
| Data in CIF |  emd_6911_validation.cif.gz emd_6911_validation.cif.gz | 6.6 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6911 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6911 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6911 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6911 | HTTPS FTP |
-Related structure data
| Related structure data |  5zbgMC  6856C  5yx9C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6911.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6911.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.37 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
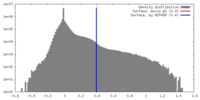
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : hTRPC3 homo-tetramer
| Entire | Name: hTRPC3 homo-tetramer |
|---|---|
| Components |
|
-Supramolecule #1: hTRPC3 homo-tetramer
| Supramolecule | Name: hTRPC3 homo-tetramer / type: organelle_or_cellular_component / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Short transient receptor potential channel 3
| Macromolecule | Name: Short transient receptor potential channel 3 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 97.465836 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MEGSPSLRRM TVMREKGRRQ AVRGPAFMFN DRGTSLTAEE ERFLDAAEYG NIPVVRKMLE ESKTLNVNCV DYMGQNALQL AVGNEHLEV TELLLKKENL ARIGDALLLA ISKGYVRIVE AILNHPGFAA SKRLTLSPCE QELQDDDFYA YDEDGTRFSP D ITPIILAA ...String: MEGSPSLRRM TVMREKGRRQ AVRGPAFMFN DRGTSLTAEE ERFLDAAEYG NIPVVRKMLE ESKTLNVNCV DYMGQNALQL AVGNEHLEV TELLLKKENL ARIGDALLLA ISKGYVRIVE AILNHPGFAA SKRLTLSPCE QELQDDDFYA YDEDGTRFSP D ITPIILAA HCQKYEVVHM LLMKGARIER PHDYFCKCGD CMEKQRHDSF SHSRSRINAY KGLASPAYLS LSSEDPVLTA LE LSNELAK LANIEKEFKN DYRKLSMQCK DFVVGVLDLC RDSEEVEAIL NGDLESAEPL EVHRHKASLS RVKLAIKYEV KKF VAHPNC QQQLLTIWYE NLSGLREQTI AIKCLVVLVV ALGLPFLAIG YWIAPCSRLG KILRSPFMKF VAHAASFIIF LGLL VFNAS DRFEGITTLP NITVTDYPKQ IFRVKTTQFT WTEMLIMVWV LGMMWSECKE LWLEGPREYI LQLWNVLDFG MLSIF IAAF TARFLAFLQA TKAQQYVDSY VQESDLSEVT LPPEIQYFTY ARDKWLPSDP QIISEGLYAI AVVLSFSRIA YILPAN ESF GPLQISLGRT VKDIFKFMVL FIMVFFAFMI GMFILYSYYL GAKVNAAFTT VEESFKTLFW SIFGLSEVTS VVLKYDH KF IENIGYVLYG IYNVTMVVVL LNMLIAMINS SYQEIEDDSD VEWKFARSKL WLSYFDDGKT LPPPFSLVPS PKSFVYFI M RIVNFPKCRR RRLQKDIEMG MGNSKSRLNL FTQSNSRVFE SHSFNSILNQ PTRYQQIMKR LIKRYVLKAQ VDKENDEVN EGELKEIKQD ISSLRYELLE DKSQATEELA ILIHKLSEKL NPSMLRCE UniProtKB: UNIPROTKB: Q13507 |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C4 (4 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 4.36 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC / Number images used: 22220 |
| Initial angle assignment | Type: OTHER |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)