+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6878 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of a nucleolar pre-60S ribosome (Rpf1-TAP) | |||||||||
 Map data Map data | Unmasked cryo-EM map | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ribosome / pre-60S / pre-ribosome / protein-RNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsnoRNA release from pre-rRNA / nuclear division / exonucleolytic trimming to generate mature 5'-end of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / endonucleolytic cleavage in ITS1 upstream of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / PeBoW complex / rRNA primary transcript binding / ATP-dependent activity, acting on RNA / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / cytosolic large ribosomal subunit assembly / maturation of 5.8S rRNA ...snoRNA release from pre-rRNA / nuclear division / exonucleolytic trimming to generate mature 5'-end of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / endonucleolytic cleavage in ITS1 upstream of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / PeBoW complex / rRNA primary transcript binding / ATP-dependent activity, acting on RNA / maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / cytosolic large ribosomal subunit assembly / maturation of 5.8S rRNA / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / preribosome, small subunit precursor / proteasome binding / Major pathway of rRNA processing in the nucleolus and cytosol / SRP-dependent cotranslational protein targeting to membrane / ribosomal large subunit binding / GTP hydrolysis and joining of the 60S ribosomal subunit / preribosome, large subunit precursor / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / L13a-mediated translational silencing of Ceruloplasmin expression / ribosomal large subunit export from nucleus / translational elongation / 90S preribosome / ribonucleoprotein complex binding / ribosomal subunit export from nucleus / maturation of LSU-rRNA / translation initiation factor activity / nuclear periphery / proteasome complex / assembly of large subunit precursor of preribosome / cytosolic ribosome assembly / ribosomal large subunit biogenesis / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / macroautophagy / protein catabolic process / maturation of SSU-rRNA / small-subunit processome / maintenance of translational fidelity / rRNA processing / nuclear envelope / ribosomal small subunit biogenesis / ribosomal large subunit assembly / 5S rRNA binding / large ribosomal subunit rRNA binding / protein-macromolecule adaptor activity / cytosolic large ribosomal subunit / cytoplasmic translation / RNA helicase activity / negative regulation of translation / rRNA binding / structural constituent of ribosome / RNA helicase / ribosome / translation / mRNA binding / nucleolus / ATP hydrolysis activity / RNA binding / zinc ion binding / nucleoplasm / ATP binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.65 Å | |||||||||
 Authors Authors | Zhou D / Zhu X / Ye K | |||||||||
 Citation Citation |  Journal: Protein Cell / Year: 2019 Journal: Protein Cell / Year: 2019Title: Cryo-EM structure of an early precursor of large ribosomal subunit reveals a half-assembled intermediate. Authors: Dejian Zhou / Xing Zhu / Sanduo Zheng / Dan Tan / Meng-Qiu Dong / Keqiong Ye /  Abstract: Assembly of eukaryotic ribosome is a complicated and dynamic process that involves a series of intermediates. It is unknown how the highly intertwined structure of 60S large ribosomal subunits is ...Assembly of eukaryotic ribosome is a complicated and dynamic process that involves a series of intermediates. It is unknown how the highly intertwined structure of 60S large ribosomal subunits is established. Here, we report the structure of an early nucleolar pre-60S ribosome determined by cryo-electron microscopy at 3.7 Å resolution, revealing a half-assembled subunit. Domains I, II and VI of 25S/5.8S rRNA pack tightly into a native-like substructure, but domains III, IV and V are not assembled. The structure contains 12 assembly factors and 19 ribosomal proteins, many of which are required for early processing of large subunit rRNA. The Brx1-Ebp2 complex would interfere with the assembly of domains IV and V. Rpf1, Mak16, Nsa1 and Rrp1 form a cluster that consolidates the joining of domains I and II. Our structure reveals a key intermediate on the path to establishing the global architecture of 60S subunits. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6878.map.gz emd_6878.map.gz | 166.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6878-v30.xml emd-6878-v30.xml emd-6878.xml emd-6878.xml | 54.9 KB 54.9 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_6878_fsc.xml emd_6878_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_6878.png emd_6878.png | 56.7 KB | ||
| Filedesc metadata |  emd-6878.cif.gz emd-6878.cif.gz | 13.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6878 http://ftp.pdbj.org/pub/emdb/structures/EMD-6878 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6878 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6878 | HTTPS FTP |
-Related structure data
| Related structure data |  5z3gMC  5z1gC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6878.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6878.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Unmasked cryo-EM map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.38 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
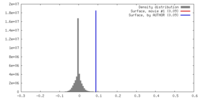
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : Rpf1-TAP pre-60S
+Supramolecule #1: Rpf1-TAP pre-60S
+Macromolecule #1: 25S rRNA
+Macromolecule #2: 5.8S rRNA
+Macromolecule #3: ITS2 RNA
+Macromolecule #4: Ribosome biogenesis protein RLP7
+Macromolecule #5: Ribosome biogenesis protein 15
+Macromolecule #6: 60S ribosomal protein L3
+Macromolecule #7: 60S ribosomal protein L4-A
+Macromolecule #8: Proteasome-interacting protein CIC1
+Macromolecule #9: 60S ribosomal protein L6-A
+Macromolecule #10: 60S ribosomal protein L7-A
+Macromolecule #11: 60S ribosomal protein L8-A
+Macromolecule #12: 60S ribosomal protein L9-A
+Macromolecule #13: Nucleolar protein 7
+Macromolecule #14: Eukaryotic translation initiation factor 6
+Macromolecule #15: Ribosome production factor 1
+Macromolecule #16: 60S ribosomal protein L13-A
+Macromolecule #17: 60S ribosomal protein L14-A
+Macromolecule #18: 60S ribosomal protein L15-A
+Macromolecule #19: 60S ribosomal protein L16-A
+Macromolecule #20: 60S ribosomal protein L17-A
+Macromolecule #21: 60S ribosomal protein L18-A
+Macromolecule #22: Ribosome biogenesis protein NSA1
+Macromolecule #23: 60S ribosomal protein L20-A
+Macromolecule #24: Ribosomal RNA-processing protein 1
+Macromolecule #25: ATP-dependent RNA helicase HAS1
+Macromolecule #26: Protein MAK16
+Macromolecule #27: Ribosome biogenesis protein BRX1
+Macromolecule #28: rRNA-processing protein EBP2
+Macromolecule #29: 60S ribosomal protein L26-A
+Macromolecule #30: Unassigned
+Macromolecule #31: 60S ribosomal protein L32
+Macromolecule #32: 60S ribosomal protein L33-A
+Macromolecule #33: 60S ribosomal protein L35-A
+Macromolecule #34: 60S ribosomal protein L36-A
+Macromolecule #35: 60S ribosomal protein L37-A
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 Component:
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Material: COPPER / Mesh: 400 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot for 1-3 seconds before plunging. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Number grids imaged: 1 / Number real images: 1983 / Average exposure time: 1.5 sec. / Average electron dose: 30.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | real space refinement |
|---|---|
| Refinement | Space: REAL |
| Output model |  PDB-5z3g: |
 Movie
Movie Controller
Controller





















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)