+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Ovorubin from the golden apple snail (Pomacea canaliculata) | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | ovorubin / golden apple snail / Pomacea canaliculata / LIPID BINDING PROTEIN | |||||||||
| Function / homology | Uncharacterized protein / Perivitellin protein / Perivitellin ovorubin-2 / Perivitellin ovorubin-1 Function and homology information Function and homology information | |||||||||
| Biological species |  Pomacea canaliculata (invertebrata) Pomacea canaliculata (invertebrata) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.12 Å | |||||||||
 Authors Authors | Wangkanont K / Saw W-G / Tran BN / Wilasluck P | |||||||||
| Funding support |  Thailand, 1 items Thailand, 1 items
| |||||||||
 Citation Citation |  Journal: Protein Sci / Year: 2026 Journal: Protein Sci / Year: 2026Title: Structure of the chromoprotein ovorubin from the golden apple snail (Pomacea canaliculata). Authors: Patcharin Wilasluck / Wuan-Geok Saw / Bich Ngoc Tran / Grzegorz Sabat / Orion Shih / Kowit Hengphasatporn / Yasuteru Shigeta / Nawaporn Vinayavekhin / Kittikhun Wangkanont /      Abstract: The golden apple snail (Pomacea canaliculata), an invasive gastropod, produces distinctly bright pink egg masses. The astaxanthin-binding ovorubin (P. canaliculata ovorubin [PcOvo]) is a 300-kDa ...The golden apple snail (Pomacea canaliculata), an invasive gastropod, produces distinctly bright pink egg masses. The astaxanthin-binding ovorubin (P. canaliculata ovorubin [PcOvo]) is a 300-kDa glycoprotein responsible for the egg coloration. Here, we determine the three-dimensional structure of PcOvo using cryo-electron microscopy (cryo-EM). PcOvo is a heterodecameric protein consisting of two copies of each PcOvo1-5 subunit. The subunits have similar repeated ferredoxin-like structures. N-linked glycosylation sites are identified. Solution x-ray scattering data support the overall architecture of the complex. PcOvo3 and PcOvo5 have hydrophobic pockets that likely bind various hydrophobic compounds. The cryo-EM map and binding experiments suggest that PcOvo5 binds astaxanthin, which is responsible for the pink color of the protein. Our structure provides molecular insights into the nature of the gastropod egg coloration and lays a foundation for further investigation of ovorubin biology and evolution. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_63984.map.gz emd_63984.map.gz | 62.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-63984-v30.xml emd-63984-v30.xml emd-63984.xml emd-63984.xml | 32.6 KB 32.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_63984_fsc.xml emd_63984_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_63984.png emd_63984.png | 162.7 KB | ||
| Masks |  emd_63984_msk_1.map emd_63984_msk_1.map | 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-63984.cif.gz emd-63984.cif.gz | 7.6 KB | ||
| Others |  emd_63984_additional_1.map.gz emd_63984_additional_1.map.gz emd_63984_half_map_1.map.gz emd_63984_half_map_1.map.gz emd_63984_half_map_2.map.gz emd_63984_half_map_2.map.gz | 109.2 MB 116.1 MB 116.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-63984 http://ftp.pdbj.org/pub/emdb/structures/EMD-63984 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63984 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-63984 | HTTPS FTP |
-Related structure data
| Related structure data |  9uajMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_63984.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_63984.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.58 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_63984_msk_1.map emd_63984_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: Sharpen map from deepEMhancer
| File | emd_63984_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Sharpen map from deepEMhancer | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_63984_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_63984_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Ovorubin
| Entire | Name: Ovorubin |
|---|---|
| Components |
|
-Supramolecule #1: Ovorubin
| Supramolecule | Name: Ovorubin / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 |
|---|---|
| Source (natural) | Organism:  Pomacea canaliculata (invertebrata) Pomacea canaliculata (invertebrata) |
| Molecular weight | Theoretical: 226 KDa |
-Macromolecule #1: PcOvo5
| Macromolecule | Name: PcOvo5 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pomacea canaliculata (invertebrata) Pomacea canaliculata (invertebrata) |
| Molecular weight | Theoretical: 22.841211 KDa |
| Sequence | String: MLVATLTLVA LSAVFTNVYG SDQYTILDVY KASNVSVEDY KDLLKDLDVV HSFKVLGSSR VIFVVKMRED SYEKLSKINL PGDVYSIPA GDLSDKMQSV GVEWKRWDDL PDANLTLFER TLELKGEPLE GLASHMKAFG EKVSHVMELY PNKGFYLLGR T PPKAFVIV ...String: MLVATLTLVA LSAVFTNVYG SDQYTILDVY KASNVSVEDY KDLLKDLDVV HSFKVLGSSR VIFVVKMRED SYEKLSKINL PGDVYSIPA GDLSDKMQSV GVEWKRWDDL PDANLTLFER TLELKGEPLE GLASHMKAFG EKVSHVMELY PNKGFYLLGR T PPKAFVIV SLPFRCRQVR YGSDFALNYL NGPGDSSTKV EFVAKA UniProtKB: Uncharacterized protein |
-Macromolecule #2: PcOvo3s
| Macromolecule | Name: PcOvo3s / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pomacea canaliculata (invertebrata) Pomacea canaliculata (invertebrata) |
| Molecular weight | Theoretical: 22.252473 KDa |
| Sequence | String: MYALAIALLA FSTFVSNAFA NKEYLLLDIR DATTSEIISA LRDVEIELKV KAKGIARHLI VVKQNDANLQ KLGEIDIPGR SCSTPVEDL DNLMEDIGIS WPRNELTNVN VTLFERTLDL KDKTMEQFWS EAKAYGQLVK PVLSSFTYRA FKANGAYPPK V YFFVNLPR ...String: MYALAIALLA FSTFVSNAFA NKEYLLLDIR DATTSEIISA LRDVEIELKV KAKGIARHLI VVKQNDANLQ KLGEIDIPGR SCSTPVEDL DNLMEDIGIS WPRNELTNVN VTLFERTLDL KDKTMEQFWS EAKAYGQLVK PVLSSFTYRA FKANGAYPPK V YFFVNLPR ENLNDASSKG IDIFGGPGKA RTTVQYVTKL S UniProtKB: Uncharacterized protein |
-Macromolecule #3: Perivitellin ovorubin-1
| Macromolecule | Name: Perivitellin ovorubin-1 / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pomacea canaliculata (invertebrata) Pomacea canaliculata (invertebrata) |
| Molecular weight | Theoretical: 22.408697 KDa |
| Sequence | String: MFVATSLLLA IATLVVRASP HNYLIMDIEP PKSVSERDIL NLLSPLQVKH SFRVTGSTRL LIVIRLDAQS YEKLDEITVP GKVEVIPAV NMADTMERCG VSWPRVELTD DNVTLFESES TLTDVTKEQL KAMLIGYGEH MSGLLQAHRF EYYQAAGATP H RHFVFVNS ...String: MFVATSLLLA IATLVVRASP HNYLIMDIEP PKSVSERDIL NLLSPLQVKH SFRVTGSTRL LIVIRLDAQS YEKLDEITVP GKVEVIPAV NMADTMERCG VSWPRVELTD DNVTLFESES TLTDVTKEQL KAMLIGYGEH MSGLLQAHRF EYYQAAGATP H RHFVFVNS VPDEIEVFGR EGVDIWGGPG EFVVKPQYVT RI UniProtKB: Perivitellin ovorubin-1 |
-Macromolecule #4: Perivitellin protein
| Macromolecule | Name: Perivitellin protein / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pomacea canaliculata (invertebrata) Pomacea canaliculata (invertebrata) |
| Molecular weight | Theoretical: 22.467189 KDa |
| Sequence | String: MTAILALLLA MSILLPSGAV EDVQDLVFAE WDKGSSHEHA CSALRNSSVI EKGLTVKEVG TSKFAAVLSE PILARLKFHG LVEAVPVVE VGTVMKRLNV SIPPAQDISD NNLTLIKMSP KLKGQTLQQI DAELRYLGEY MNTVLQKCSH RVYISKGTFP P KIYVFLNM ...String: MTAILALLLA MSILLPSGAV EDVQDLVFAE WDKGSSHEHA CSALRNSSVI EKGLTVKEVG TSKFAAVLSE PILARLKFHG LVEAVPVVE VGTVMKRLNV SIPPAQDISD NNLTLIKMSP KLKGQTLQQI DAELRYLGEY MNTVLQKCSH RVYISKGTFP P KIYVFLNM PLDQIRQFYP SLDIFGGPSS TKNEISYVQI LILRN UniProtKB: Perivitellin protein |
-Macromolecule #5: Perivitellin ovorubin-2
| Macromolecule | Name: Perivitellin ovorubin-2 / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Pomacea canaliculata (invertebrata) Pomacea canaliculata (invertebrata) |
| Molecular weight | Theoretical: 23.827533 KDa |
| Sequence | String: MAEHRIPVLL LLVVVAASSL AQQIPLQKHY IIYEVRNIEK TPEEVKEEMK DTDILYSFKA LGAPSYHIVV EVNPRNMRKL EEVELKGKI RMVPVVNMVD VAETLGVSWP RSGARLLDVN LTLIERTLNQ EGLTSQESEA HLKGFMEELK DRLQQYNYQA F FTIGASPP ...String: MAEHRIPVLL LLVVVAASSL AQQIPLQKHY IIYEVRNIEK TPEEVKEEMK DTDILYSFKA LGAPSYHIVV EVNPRNMRKL EEVELKGKI RMVPVVNMVD VAETLGVSWP RSGARLLDVN LTLIERTLNQ EGLTSQESEA HLKGFMEELK DRLQQYNYQA F FTIGASPP KMYIYINIPY EEVDKFACIG INQFGGPAAV NTTVSFISSF PK UniProtKB: Perivitellin ovorubin-2 |
-Macromolecule #8: (Z)-docos-13-enoic acid
| Macromolecule | Name: (Z)-docos-13-enoic acid / type: ligand / ID: 8 / Number of copies: 4 / Formula: 08O |
|---|---|
| Molecular weight | Theoretical: 338.568 Da |
| Chemical component information | 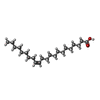 ChemComp-08O: |
-Macromolecule #9: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 9 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #10: water
| Macromolecule | Name: water / type: ligand / ID: 10 / Number of copies: 319 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 9.9 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Component - Concentration: 50.0 mM / Component - Name: Tris |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.5 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: TFS Selectris X / Energy filter - Slit width: 10 eV |
| Software | Name: EPU (ver. 3.3) |
| Image recording | Film or detector model: FEI FALCON IV (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Number grids imaged: 1 / Number real images: 6418 / Average exposure time: 2.0 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.4000000000000001 µm / Nominal defocus min: 0.4 µm / Nominal magnification: 215000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Software | Name:  Coot (ver. 0.9.8.93) Coot (ver. 0.9.8.93) | |||||||||||||||||||||||||||||||||
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 51.4 | |||||||||||||||||||||||||||||||||
| Output model |  PDB-9uaj: |
 Movie
Movie Controller
Controller




 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)
























































