Yorodumi
Yorodumi+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-5976 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structures of yeast 80S ribosome-tRNA complexes in the rotated and non-rotated conformations (Class II - 1 tRNA in rotated conformation) | |||||||||
 Map data Map data | Reconstruction of a yeast 80S ribosome in the rotated state with 1 tRNA bound. (Class II) | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | 80S ribosome / Kozak sequence / translation | |||||||||
| Function / homology |  Function and homology information Function and homology informationribophagy / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, LSU-rRNA,5S) / regulation of amino acid metabolic process / negative regulation of glucose mediated signaling pathway / translational readthrough / positive regulation of translational fidelity / : / RMTs methylate histone arginines / Protein methylation / mTORC1-mediated signalling ...ribophagy / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, LSU-rRNA,5S) / regulation of amino acid metabolic process / negative regulation of glucose mediated signaling pathway / translational readthrough / positive regulation of translational fidelity / : / RMTs methylate histone arginines / Protein methylation / mTORC1-mediated signalling / Protein hydroxylation / positive regulation of protein kinase activity / ribosome-associated ubiquitin-dependent protein catabolic process / pre-mRNA 5'-splice site binding / GDP-dissociation inhibitor activity / cytosolic large ribosomal subunit assembly / positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / nonfunctional rRNA decay / Formation of the ternary complex, and subsequently, the 43S complex / Translation initiation complex formation / response to cycloheximide / Ribosomal scanning and start codon recognition / preribosome, small subunit precursor / cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / Major pathway of rRNA processing in the nucleolus and cytosol / mRNA destabilization / SRP-dependent cotranslational protein targeting to membrane / GTP hydrolysis and joining of the 60S ribosomal subunit / negative regulation of mRNA splicing, via spliceosome / preribosome, large subunit precursor / Formation of a pool of free 40S subunits / Nonsense Mediated Decay (NMD) independent of the Exon Junction Complex (EJC) / Nonsense Mediated Decay (NMD) enhanced by the Exon Junction Complex (EJC) / L13a-mediated translational silencing of Ceruloplasmin expression / negative regulation of translational frameshifting / translational elongation / G-protein alpha-subunit binding / ribosomal large subunit export from nucleus / 90S preribosome / Ub-specific processing proteases / endonucleolytic cleavage to generate mature 3'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / ribosomal subunit export from nucleus / regulation of translational fidelity / translational termination / protein-RNA complex assembly / maturation of LSU-rRNA / endonucleolytic cleavage in ITS1 to separate SSU-rRNA from 5.8S rRNA and LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / translation regulator activity / ribosomal small subunit export from nucleus / DNA-(apurinic or apyrimidinic site) endonuclease activity / rescue of stalled cytosolic ribosome / cellular response to amino acid starvation / protein kinase C binding / ribosomal large subunit biogenesis / ribosome assembly / maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / macroautophagy / maturation of SSU-rRNA / small-subunit processome / translational initiation / maintenance of translational fidelity / modification-dependent protein catabolic process / protein tag activity / cytoplasmic stress granule / rRNA processing / ribosomal small subunit assembly / ribosome biogenesis / ribosome binding / ribosomal small subunit biogenesis / 5S rRNA binding / small ribosomal subunit / ribosomal large subunit assembly / small ribosomal subunit rRNA binding / cytosolic small ribosomal subunit / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / negative regulation of translation / rRNA binding / structural constituent of ribosome / protein ubiquitination / ribosome / translation / G protein-coupled receptor signaling pathway / negative regulation of gene expression / response to antibiotic / mRNA binding / ubiquitin protein ligase binding / nucleolus / mitochondrion / RNA binding / zinc ion binding / nucleoplasm / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.2 Å | |||||||||
 Authors Authors | Svidritskiy E / Brilot AF / Koh CS / Grigorieff N / Korostelev AA | |||||||||
 Citation Citation |  Journal: Structure / Year: 2014 Journal: Structure / Year: 2014Title: Structures of yeast 80S ribosome-tRNA complexes in the rotated and nonrotated conformations. Authors: Egor Svidritskiy / Axel F Brilot / Cha San Koh / Nikolaus Grigorieff / Andrei A Korostelev /  Abstract: The structural understanding of eukaryotic translation lags behind that of translation on bacterial ribosomes. Here, we present two subnanometer resolution structures of S. cerevisiae 80S ribosome ...The structural understanding of eukaryotic translation lags behind that of translation on bacterial ribosomes. Here, we present two subnanometer resolution structures of S. cerevisiae 80S ribosome complexes formed with either one or two tRNAs and bound in response to an mRNA fragment containing the Kozak consensus sequence. The ribosomes adopt two globally different conformations that are related to each other by the rotation of the small subunit. Comparison with bacterial ribosome complexes reveals that the global structures and modes of intersubunit rotation of the yeast ribosome differ significantly from those in the bacterial counterpart, most notably in the regions involving the tRNA, small ribosomal subunit, and conserved helix 69 of the large ribosomal subunit. The structures provide insight into ribosome dynamics implicated in tRNA translocation and help elucidate the role of the Kozak fragment in positioning an open reading frame during translation initiation in eukaryotes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_5976.map.gz emd_5976.map.gz | 161.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-5976-v30.xml emd-5976-v30.xml emd-5976.xml emd-5976.xml | 16.1 KB 16.1 KB | Display Display |  EMDB header EMDB header |
| Images |  400_5976.gif 400_5976.gif 80_5976.gif 80_5976.gif | 60.9 KB 4.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-5976 http://ftp.pdbj.org/pub/emdb/structures/EMD-5976 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5976 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-5976 | HTTPS FTP |
-Related structure data
| Related structure data |  3j77MC  5977C  3j78C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10016 (Title: Yeast 80S Ribosome - tRNA- Kozak mRNA complexes, Frealign Input Particle Stack EMPIAR-10016 (Title: Yeast 80S Ribosome - tRNA- Kozak mRNA complexes, Frealign Input Particle StackData size: 42.0 Data #1: Frealign input particle stack [picked particles - multiframe - processed]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_5976.map.gz / Format: CCP4 / Size: 173.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_5976.map.gz / Format: CCP4 / Size: 173.8 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of a yeast 80S ribosome in the rotated state with 1 tRNA bound. (Class II) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
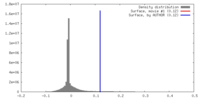
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 80S ribosome bound to mRNA containing Kozak sequence and to one tRNA
| Entire | Name: 80S ribosome bound to mRNA containing Kozak sequence and to one tRNA |
|---|---|
| Components |
|
-Supramolecule #1000: 80S ribosome bound to mRNA containing Kozak sequence and to one tRNA
| Supramolecule | Name: 80S ribosome bound to mRNA containing Kozak sequence and to one tRNA type: sample / ID: 1000 / Details: Sample was monodisperse. / Number unique components: 3 |
|---|---|
| Molecular weight | Experimental: 3.5 MDa |
-Supramolecule #1: 80S ribosome
| Supramolecule | Name: 80S ribosome / type: complex / ID: 1 / Recombinant expression: No / Database: NCBI / Ribosome-details: ribosome-eukaryote: ALL |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Experimental: 3.5 MDa |
-Macromolecule #1: mRNA
| Macromolecule | Name: mRNA / type: rna / ID: 1 / Classification: OTHER / Structure: SINGLE STRANDED / Synthetic?: Yes |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 5 KDa |
| Sequence | String: AAAAAUGUAA AAAA |
-Macromolecule #2: transfer RNA
| Macromolecule | Name: transfer RNA / type: rna / ID: 2 / Name.synonym: tRNA / Details: tRNA fmet / Classification: TRANSFER / Structure: OTHER / Synthetic?: No |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 25 KDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.2 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 20 mM Tris-HCl, 50 mM NH4Cl, 20 mM MgCl2, 0.3 U/uL RNasin |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Instrument: FEI VITROBOT MARK II |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Date | Jan 2, 2013 |
| Image recording | Category: CCD / Film or detector model: FEI FALCON I (4k x 4k) / Digitization - Sampling interval: 14 µm / Number real images: 4754 / Average electron dose: 30 e/Å2 / Bits/pixel: 16 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated magnification: 133333 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal defocus max: 4.844 µm / Nominal defocus min: 1.159 µm / Nominal magnification: 133333 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Details: CTFFIND3, FREALIGN per micrograph |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 6.2 Å / Resolution method: OTHER / Software - Name: EMAN2, IMAGIC, FREALIGN, RSAMPLE, CTFFIND3 / Number images used: 25136 |
-Atomic model buiding 1
| Initial model | PDB ID:  3u5b |
|---|---|
| Software | Name: Chimera, CNS |
| Details | 3U5B, 3U5C, 3U5D, and 3U5E were combined prior to fitting. tRNA and mRNA were modeled using individual tRNA and mRNA from the crystal structure (4GD1) of the hybrid-state 70S ribosome containing P/E tRNA. The structure of rpL1 was obtained by homology modeling from PDB ID 3J3B. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: cross-correlation |
| Output model |  PDB-3j77: |
-Atomic model buiding 2
| Initial model | PDB ID:  3u5c |
|---|---|
| Software | Name: Chimera, CNS |
| Details | 3U5B, 3U5C, 3U5D, and 3U5E were combined prior to fitting. tRNA and mRNA were modeled using individual tRNA and mRNA from the crystal structure (4GD1) of the hybrid-state 70S ribosome containing P/E tRNA. The structure of rpL1 was obtained by homology modeling from PDB ID 3J3B. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: cross-correlation |
| Output model |  PDB-3j77: |
-Atomic model buiding 3
| Initial model | PDB ID:  3u5d |
|---|---|
| Software | Name: Chimera, CNS |
| Details | 3U5B, 3U5C, 3U5D, and 3U5E were combined prior to fitting. tRNA and mRNA were modeled using individual tRNA and mRNA from the crystal structure (4GD1) of the hybrid-state 70S ribosome containing P/E tRNA. The structure of rpL1 was obtained by homology modeling from PDB ID 3J3B. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: cross-correlation |
| Output model |  PDB-3j77: |
-Atomic model buiding 4
| Initial model | PDB ID:  3u5e |
|---|---|
| Software | Name: Chimera, CNS |
| Details | 3U5B, 3U5C, 3U5D, and 3U5E were combined prior to fitting. tRNA and mRNA were modeled using individual tRNA and mRNA from the crystal structure (4GD1) of the hybrid-state 70S ribosome containing P/E tRNA. The structure of rpL1 was obtained by homology modeling from PDB ID 3J3B. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: cross-correlation |
| Output model |  PDB-3j77: |
-Atomic model buiding 5
| Initial model | PDB ID:  4gd1 Chain - #0 - Chain ID: V / Chain - #1 - Chain ID: X |
|---|---|
| Software | Name: Chimera, CNS |
| Details | 3U5B, 3U5C, 3U5D, and 3U5E were combined prior to fitting. tRNA and mRNA were modeled using individual tRNA and mRNA from the crystal structure (4GD1) of the hybrid-state 70S ribosome containing P/E tRNA. The structure of rpL1 was obtained by homology modeling from PDB ID 3J3B. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: cross-correlation |
| Output model |  PDB-3j77: |
-Atomic model buiding 6
| Initial model | PDB ID:  3j3b |
|---|---|
| Software | Name: Chimera, CNS |
| Details | 3U5B, 3U5C, 3U5D, and 3U5E were combined prior to fitting. tRNA and mRNA were modeled using individual tRNA and mRNA from the crystal structure (4GD1) of the hybrid-state 70S ribosome containing P/E tRNA. The structure of rpL1 was obtained by homology modeling from PDB ID 3J3B. |
| Refinement | Space: REAL / Protocol: RIGID BODY FIT / Target criteria: cross-correlation |
| Output model |  PDB-3j77: |
 Movie
Movie Controller
Controller


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)