[English] 日本語
 Yorodumi
Yorodumi- EMDB-43480: Structure of a synthetic antibody in complex with a class I MHC p... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of a synthetic antibody in complex with a class I MHC presenting a hapten-peptide conjugate | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Complex / Synthetic antibody / MHC class I / covalently modified peptide / IMMUNE SYSTEM | |||||||||
| Function / homology |  Function and homology information Function and homology informationresponse to mineralocorticoid / GMP binding / forebrain astrocyte development / LRR domain binding / regulation of synaptic transmission, GABAergic / negative regulation of epithelial cell differentiation / response to isolation stress / response to gravity / epithelial tube branching involved in lung morphogenesis / type I pneumocyte differentiation ...response to mineralocorticoid / GMP binding / forebrain astrocyte development / LRR domain binding / regulation of synaptic transmission, GABAergic / negative regulation of epithelial cell differentiation / response to isolation stress / response to gravity / epithelial tube branching involved in lung morphogenesis / type I pneumocyte differentiation / Rac protein signal transduction / myoblast proliferation / Signaling by RAS GAP mutants / Signaling by RAS GTPase mutants / Activation of RAS in B cells / RAS signaling downstream of NF1 loss-of-function variants / RUNX3 regulates p14-ARF / positive regulation of glial cell proliferation / skeletal muscle cell differentiation / SOS-mediated signalling / Activated NTRK3 signals through RAS / Activated NTRK2 signals through RAS / SHC1 events in ERBB4 signaling / cardiac muscle cell proliferation / Signalling to RAS / antigen processing and presentation of peptide antigen via MHC class I / SHC-related events triggered by IGF1R / Activated NTRK2 signals through FRS2 and FRS3 / positive regulation of Rac protein signal transduction / Estrogen-stimulated signaling through PRKCZ / SHC-mediated cascade:FGFR3 / glial cell proliferation / MET activates RAS signaling / SHC-mediated cascade:FGFR2 / SHC-mediated cascade:FGFR4 / Signaling by PDGFRA transmembrane, juxtamembrane and kinase domain mutants / Signaling by PDGFRA extracellular domain mutants / PTK6 Regulates RHO GTPases, RAS GTPase and MAP kinases / Erythropoietin activates RAS / SHC-mediated cascade:FGFR1 / Signaling by FGFR4 in disease / Signaling by CSF3 (G-CSF) / FRS-mediated FGFR3 signaling / Signaling by FLT3 ITD and TKD mutants / FRS-mediated FGFR2 signaling / FRS-mediated FGFR4 signaling / p38MAPK events / FRS-mediated FGFR1 signaling / Signaling by FGFR3 in disease / protein-membrane adaptor activity / striated muscle cell differentiation / Tie2 Signaling / Signaling by FGFR2 in disease / GRB2 events in EGFR signaling / Signaling by FLT3 fusion proteins / SHC1 events in EGFR signaling / FLT3 Signaling / Signaling by FGFR1 in disease / EGFR Transactivation by Gastrin / NCAM signaling for neurite out-growth / CD209 (DC-SIGN) signaling / GRB2 events in ERBB2 signaling / Downstream signal transduction / homeostasis of number of cells within a tissue / Insulin receptor signalling cascade / SHC1 events in ERBB2 signaling / response to glucocorticoid / Ras activation upon Ca2+ influx through NMDA receptor / Constitutive Signaling by Overexpressed ERBB2 / Signaling by phosphorylated juxtamembrane, extracellular and kinase domain KIT mutants / early endosome lumen / Nef mediated downregulation of MHC class I complex cell surface expression / DAP12 interactions / VEGFR2 mediated cell proliferation / small monomeric GTPase / Endosomal/Vacuolar pathway / T cell mediated cytotoxicity / FCERI mediated MAPK activation / Antigen Presentation: Folding, assembly and peptide loading of class I MHC / lumenal side of endoplasmic reticulum membrane / regulation of iron ion transport / cellular response to iron(III) ion / negative regulation of iron ion transport / negative regulation of forebrain neuron differentiation / antigen processing and presentation of exogenous protein antigen via MHC class Ib, TAP-dependent / liver development / ER to Golgi transport vesicle membrane / peptide antigen assembly with MHC class I protein complex / regulation of erythrocyte differentiation / female pregnancy / Signaling by ERBB2 TMD/JMD mutants / response to molecule of bacterial origin / HFE-transferrin receptor complex / RAF activation / transferrin transport / MHC class I peptide loading complex / cellular response to iron ion / Signaling by SCF-KIT / Constitutive Signaling by EGFRvIII / negative regulation of receptor-mediated endocytosis Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
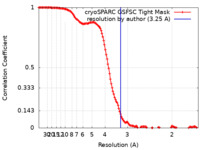
| Method | single particle reconstruction / cryo EM / Resolution: 3.25 Å | |||||||||
 Authors Authors | Maso L / Bang I / Koide S | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2024 Journal: Proc Natl Acad Sci U S A / Year: 2024Title: Molecular basis for antibody recognition of multiple drug-peptide/MHC complexes. Authors: Lorenzo Maso / Epsa Rajak / Injin Bang / Akiko Koide / Takamitsu Hattori / Benjamin G Neel / Shohei Koide /  Abstract: The HapImmune platform exploits covalent inhibitors as haptens for creating major histocompatibility complex (MHC)-presented tumor-specific neoantigens by design, combining targeted therapies with ...The HapImmune platform exploits covalent inhibitors as haptens for creating major histocompatibility complex (MHC)-presented tumor-specific neoantigens by design, combining targeted therapies with immunotherapy for the treatment of drug-resistant cancers. A HapImmune antibody, R023, recognizes multiple sotorasib-conjugated KRAS(G12C) peptides presented by different human leukocyte antigens (HLAs). This high specificity to sotorasib, coupled with broad HLA-binding capability, enables such antibodies, when reformatted as T cell engagers, to potently and selectively kill sotorasib-resistant KRAS(G12C) cancer cells expressing different HLAs upon sotorasib treatment. The loosening of HLA restriction could increase the patient population that can benefit from this therapeutic approach. To understand the molecular basis for its unconventional binding capability, we used single-particle cryogenic electron microscopy to determine the structures of R023 bound to multiple sotorasib-peptide conjugates presented by different HLAs. R023 forms a pocket for sotorasib between the V and V domains, binds HLAs in an unconventional, angled way, with V making most contacts with them, and makes few contacts with the peptide moieties. This binding mode enables the antibody to accommodate different hapten-peptide conjugates and to adjust its conformation to different HLAs presenting hapten-peptides. Deep mutational scanning validated the structures and revealed distinct levels of mutation tolerance by sotorasib- and HLA-binding residues. Together, our structural information and sequence landscape analysis reveal key features for achieving MHC-restricted recognition of multiple hapten-peptide antigens, which will inform the development of next-generation therapeutic antibodies. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_43480.map.gz emd_43480.map.gz | 203.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-43480-v30.xml emd-43480-v30.xml emd-43480.xml emd-43480.xml | 22.2 KB 22.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_43480_fsc.xml emd_43480_fsc.xml | 12.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_43480.png emd_43480.png | 95.2 KB | ||
| Filedesc metadata |  emd-43480.cif.gz emd-43480.cif.gz | 6.9 KB | ||
| Others |  emd_43480_half_map_1.map.gz emd_43480_half_map_1.map.gz emd_43480_half_map_2.map.gz emd_43480_half_map_2.map.gz | 200.6 MB 200.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-43480 http://ftp.pdbj.org/pub/emdb/structures/EMD-43480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43480 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-43480 | HTTPS FTP |
-Related structure data
| Related structure data |  8vrbMC  8vr9C  8vraC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_43480.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_43480.map.gz / Format: CCP4 / Size: 216 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.825 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_43480_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_43480_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Binary complex of Fab R023 with sotorasib-conjugated KRAS G12C pe...
+Supramolecule #1: Binary complex of Fab R023 with sotorasib-conjugated KRAS G12C pe...
+Supramolecule #2: Fab R023
+Supramolecule #3: class I MHC, having HLA-A*11:01, with Beta-2-microglobulin
+Supramolecule #4: sotorasib-conjugated KRAS G12C peptide (7-16)
+Macromolecule #1: HLA-A antigen
+Macromolecule #2: Beta-2-microglobulin
+Macromolecule #3: GTPase KRas, N-terminally processed
+Macromolecule #4: R023 Fab light chain
+Macromolecule #5: R023 Fab heavy chain
+Macromolecule #6: AMG 510 (bound form)
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.4 Component:
| ||||||||||||
| Grid | Model: Quantifoil R0.6/1 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 80 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.026000000000000002 kPa | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Average exposure time: 2.0 sec. / Average electron dose: 57.56 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 0.9 µm / Nominal magnification: 105000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)