+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human Type 3 IP3 Receptor - Resting State (+IP3/ATP) | ||||||||||||
 Map data Map data | IP3 Receptor - hIP3R3 Higher-Order Inhibited State - Consensus Refinement - Tetramer - Symmetry Mate 2 | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | Ion Channel / Calcium Channel / Endoplasmic Reticulum / TRANSPORT PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationDAG and IP3 signaling / inositol 1,3,4,5 tetrakisphosphate binding / sensory perception of bitter taste / inositol 1,4,5-trisphosphate-gated calcium channel activity / platelet dense tubular network membrane / sensory perception of umami taste / Effects of PIP2 hydrolysis / sensory perception of sweet taste / PLC beta mediated events / Elevation of cytosolic Ca2+ levels ...DAG and IP3 signaling / inositol 1,3,4,5 tetrakisphosphate binding / sensory perception of bitter taste / inositol 1,4,5-trisphosphate-gated calcium channel activity / platelet dense tubular network membrane / sensory perception of umami taste / Effects of PIP2 hydrolysis / sensory perception of sweet taste / PLC beta mediated events / Elevation of cytosolic Ca2+ levels / inositol 1,4,5 trisphosphate binding / inositol hexakisphosphate binding / transport vesicle membrane / CLEC7A (Dectin-1) induces NFAT activation / cytoplasmic side of endoplasmic reticulum membrane / intracellularly gated calcium channel activity / nuclear outer membrane / brush border / Role of phospholipids in phagocytosis / calcium ion homeostasis / Ion homeostasis / release of sequestered calcium ion into cytosol / FCERI mediated Ca+2 mobilization / phosphatidylinositol binding / FCGR3A-mediated IL10 synthesis / secretory granule membrane / Antigen activates B Cell Receptor (BCR) leading to generation of second messengers / VEGFR2 mediated cell proliferation / sarcoplasmic reticulum / Regulation of insulin secretion / platelet activation / response to calcium ion / memory / apical part of cell / Sensory perception of sweet, bitter, and umami (glutamate) taste / long-term synaptic potentiation / Glucagon-like Peptide-1 (GLP1) regulates insulin secretion / sensory perception of taste / positive regulation of cytosolic calcium ion concentration / Ca2+ pathway / protein homotetramerization / signaling receptor complex / G protein-coupled receptor signaling pathway / neuronal cell body / calcium ion binding / endoplasmic reticulum membrane / nucleolus / endoplasmic reticulum / zinc ion binding / nucleoplasm / ATP binding / membrane / plasma membrane / cytoplasm Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | ||||||||||||
 Authors Authors | Paknejad N / Sapuru V / Hite RK | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2023 Journal: Nat Commun / Year: 2023Title: Structural titration reveals Ca-dependent conformational landscape of the IP receptor. Authors: Navid Paknejad / Vinay Sapuru / Richard K Hite /  Abstract: Inositol 1,4,5-trisphosphate receptors (IPRs) are endoplasmic reticulum Ca channels whose biphasic dependence on cytosolic Ca gives rise to Ca oscillations that regulate fertilization, cell division ...Inositol 1,4,5-trisphosphate receptors (IPRs) are endoplasmic reticulum Ca channels whose biphasic dependence on cytosolic Ca gives rise to Ca oscillations that regulate fertilization, cell division and cell death. Despite the critical roles of IPR-mediated Ca responses, the structural underpinnings of the biphasic Ca dependence that underlies Ca oscillations are incompletely understood. Here, we collect cryo-EM images of an IPR with Ca concentrations spanning five orders of magnitude. Unbiased image analysis reveals that Ca binding does not explicitly induce conformational changes but rather biases a complex conformational landscape consisting of resting, preactivated, activated, and inhibited states. Using particle counts as a proxy for relative conformational free energy, we demonstrate that Ca binding at a high-affinity site allows IPRs to activate by escaping a low-energy resting state through an ensemble of preactivated states. At high Ca concentrations, IPRs preferentially enter an inhibited state stabilized by a second, low-affinity Ca binding site. Together, these studies provide a mechanistic basis for the biphasic Ca-dependence of IPR channel activity. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
-Related structure data
| Related structure data |  8tl9MC  8tk8C  8tkdC  8tkeC  8tkfC  8tkgC  8tkhC  8tkiC  8tlaC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_41365.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41365.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | IP3 Receptor - hIP3R3 Higher-Order Inhibited State - Consensus Refinement - Tetramer - Symmetry Mate 2 | ||||||||||||||||||||||||||||||||||||
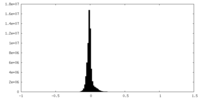
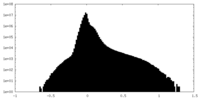
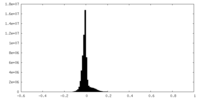
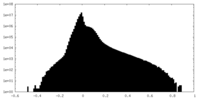
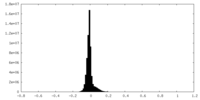
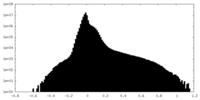
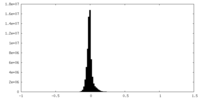
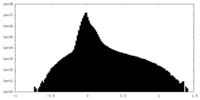
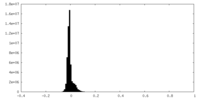
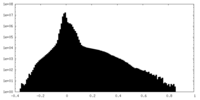
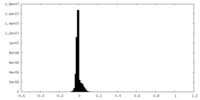
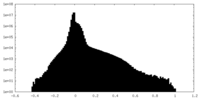
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.826 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Additional map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Half map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
+Half map: IP3 Receptor - hIP3R3 Higher-Order Inhibited State -...
- Sample components
Sample components
-Entire : Human Type 3 IP3 Receptor
| Entire | Name: Human Type 3 IP3 Receptor |
|---|---|
| Components |
|
-Supramolecule #1: Human Type 3 IP3 Receptor
| Supramolecule | Name: Human Type 3 IP3 Receptor / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 Details: In the presence of saturating IP3, ATP, and Ca2+ titrated from 1 nM to 10 um |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 1.2 MDa |
-Macromolecule #1: Inositol 1,4,5-trisphosphate receptor type 3
| Macromolecule | Name: Inositol 1,4,5-trisphosphate receptor type 3 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 304.488688 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSEMSSFLHI GDIVSLYAEG SVNGFISTLG LVDDRCVVEP AAGDLDNPPK KFRDCLFKVC PMNRYSAQKQ YWKAKQTKQD KEKIADVVL LQKLQHAAQM EQKQNDTENK KVHGDVVKYG SVIQLLHMKS NKYLTVNKRL PALLEKNAMR VTLDATGNEG S WLFIQPFW ...String: MSEMSSFLHI GDIVSLYAEG SVNGFISTLG LVDDRCVVEP AAGDLDNPPK KFRDCLFKVC PMNRYSAQKQ YWKAKQTKQD KEKIADVVL LQKLQHAAQM EQKQNDTENK KVHGDVVKYG SVIQLLHMKS NKYLTVNKRL PALLEKNAMR VTLDATGNEG S WLFIQPFW KLRSNGDNVV VGDKVILNPV NAGQPLHASN YELSDNAGCK EVNSVNCNTS WKINLFMQFR DHLEEVLKGG DV VRLFHAE QEKFLTCDEY KGKLQVFLRT TLRQSATSAT SSNALWEVEV VHHDPCRGGA GHWNGLYRFK HLATGNYLAA EEN PSYKGD ASDPKAAGMG AQGRTGRRNA GEKIKYCLVA VPHGNDIASL FELDPTTLQK TDSFVPRNSY VRLRHLCTNT WIQS TNVPI DIEEERPIRL MLGTCPTKED KEAFAIVSVP VSEIRDLDFA NDASSMLASA VEKLNEGFIS QNDRRFVIQL LEDLV FFVS DVPNNGQNVL DIMVTKPNRE RQKLMREQNI LKQVFGILKA PFREKGGEGP LVRLEELSDQ KNAPYQHMFR LCYRVL RHS QEDYRKNQEH IAKQFGMMQS QIGYDILAED TITALLHNNR KLLEKHITKT EVETFVSLVR KNREPRFLDY LSDLCVS NH IAIPVTQELI CKCVLDPKNS DILIRTELRP VKEMAQSHEY LSIEYSEEEV WLTWTDKNNE HHEKSVRQLA QEARAGNA H DENVLSYYRY QLKLFARMCL DRQYLAIDEI SQQLGVDLIF LCMADEMLPF DLRASFCHLM LHVHVDRDPQ ELVTPVKFA RLWTEIPTAI TIKDYDSNLN ASRDDKKNKF ANTMEFVEDY LNNVVSEAVP FANEEKNKLT FEVVSLAHNL IYFGFYSFSE LLRLTRTLL GIIDCVQGPP AMLQAYEDPG GKNVRRSIQG VGHMMSTMVL SRKQSVFSAP SLSAGASAAE PLDRSKFEEN E DIVVMETK LKILEILQFI LNVRLDYRIS YLLSVFKKEF VEVFPMQDSG ADGTAPAFDS TTANMNLDRI GEQAEAMFGV GK TSSMLEV DDEGGRMFLR VLIHLTMHDY APLVSGALQL LFKHFSQRQE AMHTFKQVQL LISAQDVENY KVIKSELDRL RTM VEKSEL WVDKKGSGKG EEVEAGAAKD KKERPTDEEG FLHPPGEKSS ENYQIVKGIL ERLNKMCGVG EQMRKKQQRL LKNM DAHKV MLDLLQIPYD KGDAKMMEIL RYTHQFLQKF CAGNPGNQAL LHKHLHLFLT PGLLEAETMQ HIFLNNYQLC SEISE PVLQ HFVHLLATHG RHVQYLDFLH TVIKAEGKYV KKCQDMIMTE LTNAGDDVVV FYNDKASLAH LLDMMKAARD GVEDHS PLM YHISLVDLLA ACAEGKNVYT EIKCTSLLPL EDVVSVVTHE DCITEVKMAY VNFVNHCYVD TEVEMKEIYT SNHIWTL FE NFTLDMARVC SKREKRVADP TLEKYVLSVV LDTINAFFSS PFSENSTSLQ THQTIVVQLL QSTTRLLECP WLQQQHKG S VEACIRTLAM VAKGRAILLP MDLDAHISSM LSSGASCAAA AQRNASSYKA TTRAFPRVTP TANQWDYKNI IEKLQDIIT ALEERLKPLV QAELSVLVDV LHWPELLFLE GSEAYQRCES GGFLSKLIQH TKDLMESEEK LCIKVLRTLQ QMLLKKTKYG DRGNQLRKM LLQNYLQNRK STSRGDLPDP IGTGLDPDWS AIAATQCRLD KEGATKLVCD LITSTKNEKI FQESIGLAIH L LDGGNTEI QKSFHNLMMS DKKSERFFKV LHDRMKRAQQ ETKSTVAVNM NDLGSQPHED REPVDPTTKG RVASFSIPGS SS RYSLGPS LRRGHEVSER VQSSEMGTSV LIMQPILRFL QLLCENHNRD LQNFLRCQNN KTNYNLVCET LQFLDIMCGS TTG GLGLLG LYINEDNVGL VIQTLETLTE YCQGPCHENQ TCIVTHESNG IDIITALILN DISPLCKYRM DLVLQLKDNA SKLL LALME SRHDSENAER ILISLRPQEL VDVIKKAYLQ EEERENSEVS PREVGHNIYI LALQLSRHNK QLQHLLKPVK RIQEE EAEG ISSMLSLNNK QLSQMLKSSA PAQEEEEDPL AYYENHTSQI EIVRQDRSME QIVFPVPGIC QFLTEETKHR LFTTTE QDE QGSKVSDFFD QSSFLHNEME WQRKLRSMPL IYWFSRRMTL WGSISFNLAV FINIIIAFFY PYMEGASTGV LDSPLIS LL FWILICFSIA ALFTKRYSIR PLIVALILRS IYYLGIGPTL NILGALNLTN KIVFVVSFVG NRGTFIRGYK AMVMDMEF L YHVGYILTSV LGLFAHELFY SILLFDLIYR EETLFNVIKS VTRNGRSILL TALLALILVY LFSIVGFLFL KDDFILEVD RLPNNHSTAS PLGMPHGAAA FVDTCSGDKM DCVSGLSVPE VLEEDRELDS TERACDTLLM CIVTVMNHGL RNGGGVGDIL RKPSKDESL FPARVVYDLL FFFIVIIIVL NLIFGVIIDT FADLRSEKQK KEEILKTTCF ICGLERDKFD NKTVSFEEHI K LEHNMWNY LYFIVLVRVK NKTDYTGPES YVAQMIKNKN LDWFPRMRAM SLVSNEGEGE QNEIRILQDK LNSTMKLVSH LT AQLNELK EQMTEQRKRR QRLGFVDVQN CISR UniProtKB: Inositol 1,4,5-trisphosphate-gated calcium channel ITPR3 |
-Macromolecule #2: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Macromolecule #3: D-MYO-INOSITOL-1,4,5-TRIPHOSPHATE
| Macromolecule | Name: D-MYO-INOSITOL-1,4,5-TRIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 4 / Formula: I3P |
|---|---|
| Molecular weight | Theoretical: 420.096 Da |
| Chemical component information |  ChemComp-I3P: |
-Macromolecule #4: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 4 / Number of copies: 4 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Macromolecule #5: ADENOSINE-5'-TRIPHOSPHATE
| Macromolecule | Name: ADENOSINE-5'-TRIPHOSPHATE / type: ligand / ID: 5 / Number of copies: 4 / Formula: ATP |
|---|---|
| Molecular weight | Theoretical: 507.181 Da |
| Chemical component information |  ChemComp-ATP: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 20 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 120 mM NaCl, 50 mM Tris-HCl pH 8.0, 0.02% LMNG, 2 mM dithiothreitol, 2 mM EDTA, 2 mM EGTA, 2 mM BAPTA, 2 mM HEDTA, 1 mM ATP, 500 uM fluorinated fos-choline-8, 3 mM free Mg2+, 1-10000 nM free Ca2+ |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV Details: Custom-made calcium free blotting paper (see publication for details).. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 6 / Number real images: 17733 / Average exposure time: 3.0 sec. / Average electron dose: 66.0 e/Å2 Details: Images were collected at 0.826 A/px magnification on an FEI Krios with Gatan K3 detector at 15 e-/pix/sec with 3 sec exposure (0.05 sec/frame) for a total dose of 66 e-/A2 in automated ...Details: Images were collected at 0.826 A/px magnification on an FEI Krios with Gatan K3 detector at 15 e-/pix/sec with 3 sec exposure (0.05 sec/frame) for a total dose of 66 e-/A2 in automated fashion using SerialEM. Five datasets were collected during the same session for each Ca2+ concentration on a series of grids that were prepared sequentially resulting in 637 movies at 1 nM, 2150 movies at 10 nM, 6126 movies at 100 nM, 1372 movies at 1 uM, and 3136 movies at 10 uM. A sixth dataset of 4312 movies collected at nominal 100 nM free Ca2+ from a grid prepared later in the sequence was collected as a technical replicate to assess experimental error. |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Calibrated defocus max: 4.3 µm / Calibrated defocus min: 0.7000000000000001 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 29000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: INSILICO MODEL |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.3 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC / Number images used: 88082 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: AB INITIO MODEL / Target criteria: Map to Model FSC |
|---|---|
| Output model |  PDB-8tl9: |
 Movie
Movie Controller
Controller






















































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)