+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Tetrahymena Ribozyme scaffolded Fluoride riboswitch | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Ribozyme / xrRNA / Scaffold / RNA | |||||||||
| Biological species |  Tetrahymena (eukaryote) / Tetrahymena (eukaryote) /   Thermotoga petrophila (bacteria) Thermotoga petrophila (bacteria) | |||||||||
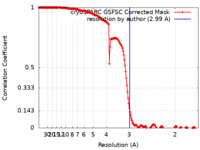
| Method | single particle reconstruction / cryo EM / Resolution: 2.99 Å | |||||||||
 Authors Authors | Langeberg CJ / Kieft JS | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res / Year: 2023 Journal: Nucleic Acids Res / Year: 2023Title: A generalizable scaffold-based approach for structure determination of RNAs by cryo-EM. Authors: Conner J Langeberg / Jeffrey S Kieft /  Abstract: Single-particle cryo-electron microscopy (cryo-EM) can reveal the structures of large and often dynamic molecules, but smaller biomolecules (≤50 kDa) remain challenging targets due to their ...Single-particle cryo-electron microscopy (cryo-EM) can reveal the structures of large and often dynamic molecules, but smaller biomolecules (≤50 kDa) remain challenging targets due to their intrinsic low signal to noise ratio. Methods to help resolve small proteins have been applied but development of similar approaches to aid in structural determination of small, structured RNA elements have lagged. Here, we present a scaffold-based approach that we used to recover maps of sub-25 kDa RNA domains to 4.5-5.0 Å. While lacking the detail of true high-resolution maps, these maps are suitable for model building and preliminary structure determination. We demonstrate this method helped faithfully recover the structure of several RNA elements of known structure, and that it promises to be generalized to other RNAs without disturbing their native fold. This approach may streamline the sample preparation process and reduce the optimization required for data collection. This first-generation scaffold approach provides a robust system to aid in RNA structure determination by cryo-EM and lays the groundwork for further scaffold optimization to achieve higher resolution. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_41312.map.gz emd_41312.map.gz | 256.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-41312-v30.xml emd-41312-v30.xml emd-41312.xml emd-41312.xml | 14.8 KB 14.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_41312_fsc.xml emd_41312_fsc.xml | 16.9 KB | Display |  FSC data file FSC data file |
| Images |  emd_41312.png emd_41312.png | 55.4 KB | ||
| Filedesc metadata |  emd-41312.cif.gz emd-41312.cif.gz | 5.2 KB | ||
| Others |  emd_41312_half_map_1.map.gz emd_41312_half_map_1.map.gz emd_41312_half_map_2.map.gz emd_41312_half_map_2.map.gz | 474.5 MB 474.5 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-41312 http://ftp.pdbj.org/pub/emdb/structures/EMD-41312 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41312 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-41312 | HTTPS FTP |
-Validation report
| Summary document |  emd_41312_validation.pdf.gz emd_41312_validation.pdf.gz | 772.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_41312_full_validation.pdf.gz emd_41312_full_validation.pdf.gz | 771.8 KB | Display | |
| Data in XML |  emd_41312_validation.xml.gz emd_41312_validation.xml.gz | 26.1 KB | Display | |
| Data in CIF |  emd_41312_validation.cif.gz emd_41312_validation.cif.gz | 34.2 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41312 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41312 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41312 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-41312 | HTTPS FTP |
-Related structure data
| Related structure data |  8tjvMC  8tjqC  8tjuC  8tjxC M: atomic model generated by this map C: citing same article ( |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_41312.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_41312.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8464 Å | ||||||||||||||||||||||||||||||||||||
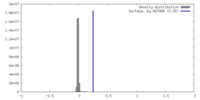
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_41312_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
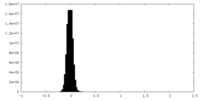
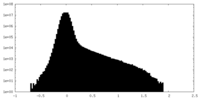
| Density Histograms |
-Half map: #1
| File | emd_41312_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
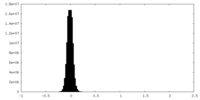
| Density Histograms |
- Sample components
Sample components
-Entire : Tetrahymena Ribozyme scaffolded Fluoride riboswitch
| Entire | Name: Tetrahymena Ribozyme scaffolded Fluoride riboswitch |
|---|---|
| Components |
|
-Supramolecule #1: Tetrahymena Ribozyme scaffolded Fluoride riboswitch
| Supramolecule | Name: Tetrahymena Ribozyme scaffolded Fluoride riboswitch / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Tetrahymena (eukaryote) Tetrahymena (eukaryote) |
-Macromolecule #1: RNA (417-MER)
| Macromolecule | Name: RNA (417-MER) / type: rna / ID: 1 / Number of copies: 1 |
|---|---|
| Source (natural) | Organism:   Thermotoga petrophila (bacteria) Thermotoga petrophila (bacteria) |
| Molecular weight | Theoretical: 134.830641 KDa |
| Sequence | String: GGGCGAUGAG GCCCGCCCAA ACUGCCCUAU AUGGAUGCAG UUCACAGACU AAAUGUCGGU CGGGGAAGAU GUAUUCUUCU CAUAAGAUA UAGUCGGACC UCUCCUUAAU GGGAGCUAGC GGAUGAAGUG AUGCAACACU GGAGCCGCUG GGAACUAAUU U GUAUGCGA ...String: GGGCGAUGAG GCCCGCCCAA ACUGCCCUAU AUGGAUGCAG UUCACAGACU AAAUGUCGGU CGGGGAAGAU GUAUUCUUCU CAUAAGAUA UAGUCGGACC UCUCCUUAAU GGGAGCUAGC GGAUGAAGUG AUGCAACACU GGAGCCGCUG GGAACUAAUU U GUAUGCGA AAGUAUAUUG AUUAGUUUUG GAGGAGGGAA AAGUUAUCAG GCAUGCACCU GGUAGCUAGU CUUUAAACCA AU AGAUUGC AUCGGUUUAA AAGGCAAGAC CGUCAAAUUG CGGGAAAGGG GUCAACAGCC GUUCAGUACC AAGUCUCAGG GGA AACUUU GAGAUGGCCU UGCAAAGGGU AUGGUAAUAA GCUGACGGAC AUGGUCCUAA CCACGCAGCC AAGUCCUAAG UAGG GCUGA UGGCCUCUAC UG |
-Macromolecule #2: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 2 / Number of copies: 30 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #3: FLUORIDE ION
| Macromolecule | Name: FLUORIDE ION / type: ligand / ID: 3 / Number of copies: 1 / Formula: F |
|---|---|
| Molecular weight | Theoretical: 18.998 Da |
| Chemical component information |  ChemComp-F: |
-Macromolecule #4: POTASSIUM ION
| Macromolecule | Name: POTASSIUM ION / type: ligand / ID: 4 / Number of copies: 1 / Formula: K |
|---|---|
| Molecular weight | Theoretical: 39.098 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: C-flat-1.2/1.3 / Material: COPPER / Mesh: 50 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 6 sec. / Pretreatment - Atmosphere: OTHER | |||||||||
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller







 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)