[English] 日本語
 Yorodumi
Yorodumi- EMDB-40222: CryoEM structure of the TnsC(1-503)-TnsD(1-318)-DNA complex in a ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of the TnsC(1-503)-TnsD(1-318)-DNA complex in a 6:2:1 stoichiometry from E. coli Tn7 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Transposon / AAA+ ATPase / Oligomer / Complex / DNA BINDING PROTEIN / DNA BINDING PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationtransposition / DNA recombination / ATP hydrolysis activity / DNA binding / ATP binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.88 Å | |||||||||
 Authors Authors | Shen Y / Guarne A | |||||||||
| Funding support |  Canada, 1 items Canada, 1 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2024 Journal: Mol Cell / Year: 2024Title: Assembly of the Tn7 targeting complex by a regulated stepwise process. Authors: Yao Shen / Shreya S Krishnan / Michael T Petassi / Mark A Hancock / Joseph E Peters / Alba Guarné /   Abstract: The Tn7 family of transposons is notable for its highly regulated integration mechanisms, including programmable RNA-guided transposition. The targeting pathways rely on dedicated target selection ...The Tn7 family of transposons is notable for its highly regulated integration mechanisms, including programmable RNA-guided transposition. The targeting pathways rely on dedicated target selection proteins from the TniQ family and the AAA+ adaptor TnsC to recruit and activate the transposase at specific target sites. Here, we report the cryoelectron microscopy (cryo-EM) structures of TnsC bound to the TniQ domain of TnsD from prototypical Tn7 and unveil key regulatory steps stemming from unique behaviors of ATP- versus ADP-bound TnsC. We show that TnsD recruits ADP-bound dimers of TnsC and acts as an exchange factor to release one protomer with exchange to ATP. This loading process explains how TnsC assembles a heptameric ring unidirectionally from the target site. This unique loading process results in functionally distinct TnsC protomers within the ring, providing a checkpoint for target immunity and explaining how insertions at programmed sites precisely occur in a specific orientation across Tn7 elements. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40222.map.gz emd_40222.map.gz | 225.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40222-v30.xml emd-40222-v30.xml emd-40222.xml emd-40222.xml | 25.5 KB 25.5 KB | Display Display |  EMDB header EMDB header |
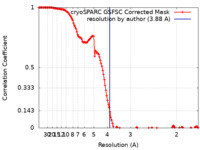
| FSC (resolution estimation) |  emd_40222_fsc.xml emd_40222_fsc.xml | 13.3 KB | Display |  FSC data file FSC data file |
| Images |  emd_40222.png emd_40222.png | 144.6 KB | ||
| Filedesc metadata |  emd-40222.cif.gz emd-40222.cif.gz | 6.8 KB | ||
| Others |  emd_40222_additional_1.map.gz emd_40222_additional_1.map.gz emd_40222_additional_2.map.gz emd_40222_additional_2.map.gz emd_40222_half_map_1.map.gz emd_40222_half_map_1.map.gz emd_40222_half_map_2.map.gz emd_40222_half_map_2.map.gz | 230 MB 124.8 MB 226.3 MB 226.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40222 http://ftp.pdbj.org/pub/emdb/structures/EMD-40222 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40222 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40222 | HTTPS FTP |
-Validation report
| Summary document |  emd_40222_validation.pdf.gz emd_40222_validation.pdf.gz | 833.3 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_40222_full_validation.pdf.gz emd_40222_full_validation.pdf.gz | 832.9 KB | Display | |
| Data in XML |  emd_40222_validation.xml.gz emd_40222_validation.xml.gz | 22.5 KB | Display | |
| Data in CIF |  emd_40222_validation.cif.gz emd_40222_validation.cif.gz | 28.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40222 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40222 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40222 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-40222 | HTTPS FTP |
-Related structure data
| Related structure data |  8glxMC  8gluC  8glwC  8vcjC  8vctC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40222.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40222.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.855 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: #1
| File | emd_40222_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #2
| File | emd_40222_additional_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_40222_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_40222_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : Complex of TnsC(1-503) bound to TnsD(1-318) and DNA with cofactor...
+Supramolecule #1: Complex of TnsC(1-503) bound to TnsD(1-318) and DNA with cofactor...
+Supramolecule #2: TnsC(1-503)
+Supramolecule #3: TnsD(1-318)
+Supramolecule #4: DNA
+Macromolecule #1: Transposon Tn7 transposition protein TnsC
+Macromolecule #2: Transposon Tn7 transposition protein TnsD
+Macromolecule #3: DNA (50-MER)
+Macromolecule #4: DNA (50-MER)
+Macromolecule #5: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #6: MAGNESIUM ION
+Macromolecule #7: ZINC ION
+Macromolecule #8: ADENOSINE-5'-DIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 Details: 20 mM Tris pH 8.0, 150 mM NaCl, 1.4 mM beta-mercaptoethanol, 5 mM MgCl2 |
| Grid | Model: C-flat-2/1 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 79.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.75 µm / Nominal defocus min: 1.25 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Source name: PDB / Chain - Initial model type: experimental model / Details: TnsC and TnsD were modelled using PDB 8GLU. |
|---|---|
| Refinement | Space: REAL / Protocol: OTHER |
| Output model |  PDB-8glx: |
 Movie
Movie Controller
Controller









 X (Sec.)
X (Sec.) Y (Row.)
Y (Row.) Z (Col.)
Z (Col.)