+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | E. coli clamp loader with open clamp | |||||||||
 Map data Map data | CLC.Beta2 Open | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | clamp loader / DNA clamp / AAA+ / ATPase / DNA BINDING PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationDNA polymerase III, clamp loader complex / Hda-beta clamp complex / bacterial-type DNA replication / replication inhibiting complex / DNA clamp loader activity / DNA polymerase III complex / replisome / regulation of DNA-templated DNA replication initiation / DNA strand elongation involved in DNA replication / DNA polymerase processivity factor activity ...DNA polymerase III, clamp loader complex / Hda-beta clamp complex / bacterial-type DNA replication / replication inhibiting complex / DNA clamp loader activity / DNA polymerase III complex / replisome / regulation of DNA-templated DNA replication initiation / DNA strand elongation involved in DNA replication / DNA polymerase processivity factor activity / error-prone translesion synthesis / 3'-5' exonuclease activity / negative regulation of DNA-templated DNA replication initiation / response to radiation / DNA-templated DNA replication / ribonucleoside triphosphate phosphatase activity / DNA-directed DNA polymerase / DNA-directed DNA polymerase activity / DNA replication / viral translational frameshifting / DNA damage response / protein homodimerization activity / ATP hydrolysis activity / DNA binding / ATP binding / metal ion binding / identical protein binding / cytosol Similarity search - Function | |||||||||
| Biological species |   | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.7 Å | |||||||||
 Authors Authors | Oakley AJ / Xu Z-Q / Dixon NE | |||||||||
| Funding support |  Australia, 1 items Australia, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural characterisation of the complete cycle of sliding clamp loading in Escherichia coli. Authors: Zhi-Qiang Xu / Slobodan Jergic / Allen T Y Lo / Alok C Pradhan / Simon H J Brown / James C Bouwer / Harshad Ghodke / Peter J Lewis / Gökhan Tolun / Aaron J Oakley / Nicholas E Dixon /  Abstract: Ring-shaped DNA sliding clamps are essential for DNA replication and genome maintenance. Clamps need to be opened and chaperoned onto DNA by clamp loader complexes (CLCs). Detailed understanding of ...Ring-shaped DNA sliding clamps are essential for DNA replication and genome maintenance. Clamps need to be opened and chaperoned onto DNA by clamp loader complexes (CLCs). Detailed understanding of the mechanisms by which CLCs open and place clamps around DNA remains incomplete. Here, we present a series of six structures of the Escherichia coli CLC bound to an open or closed clamp prior to and after binding to a primer-template DNA, representing the most significant intermediates in the clamp loading process. We show that the ATP-bound CLC first binds to a clamp, then constricts to hold onto it. The CLC then expands to open the clamp with a gap large enough for double-stranded DNA to enter. Upon binding to DNA, the CLC constricts slightly, allowing clamp closing around DNA. These structures provide critical high-resolution snapshots of clamp loading by the E. coli CLC, revealing how the molecular machine works. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_40080.map.gz emd_40080.map.gz | 140.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-40080-v30.xml emd-40080-v30.xml emd-40080.xml emd-40080.xml | 27.4 KB 27.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_40080.png emd_40080.png | 59.2 KB | ||
| Filedesc metadata |  emd-40080.cif.gz emd-40080.cif.gz | 8.1 KB | ||
| Others |  emd_40080_half_map_1.map.gz emd_40080_half_map_1.map.gz emd_40080_half_map_2.map.gz emd_40080_half_map_2.map.gz | 141.2 MB 140.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-40080 http://ftp.pdbj.org/pub/emdb/structures/EMD-40080 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40080 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-40080 | HTTPS FTP |
-Related structure data
| Related structure data |  8gizMC  8giyC  8gj0C  8gj1C  8gj2C  8gj3C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_40080.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_40080.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CLC.Beta2 Open | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.82 Å | ||||||||||||||||||||||||||||||||||||
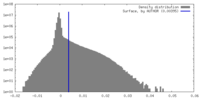
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: CLC.Beta2 Open half map 1
| File | emd_40080_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CLC.Beta2 Open half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
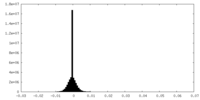
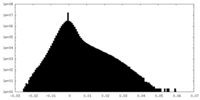
| Density Histograms |
-Half map: CLC.Beta2 Open half map 2
| File | emd_40080_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CLC.Beta2 Open half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
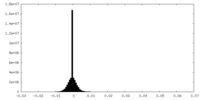
| Density Histograms |
- Sample components
Sample components
-Entire : E. coli clamp loader with open clamp
| Entire | Name: E. coli clamp loader with open clamp |
|---|---|
| Components |
|
-Supramolecule #1: E. coli clamp loader with open clamp
| Supramolecule | Name: E. coli clamp loader with open clamp / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 Details: Clamp loader complex composed of DNA polymerase III delta gamma(3) delta' chi and psi subunits. Clamp composed of DNA polymerase III beta(2) subunits. |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 331 KDa |
-Macromolecule #1: DNA polymerase III subunit delta
| Macromolecule | Name: DNA polymerase III subunit delta / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed DNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 38.745574 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MIRLYPEQLR AQLNEGLRAA YLLLGNDPLL LQESQDAVRQ VAAAQGFEEH HTFSIDPNTD WNAIFSLCQA MSLFASRQTL LLLLPENGP NAAINEQLLT LTGLLHDDLL LIVRGNKLSK AQENAAWFTA LANRSVQVTC QTPEQAQLPR WVAARAKQLN L ELDDAANQ ...String: MIRLYPEQLR AQLNEGLRAA YLLLGNDPLL LQESQDAVRQ VAAAQGFEEH HTFSIDPNTD WNAIFSLCQA MSLFASRQTL LLLLPENGP NAAINEQLLT LTGLLHDDLL LIVRGNKLSK AQENAAWFTA LANRSVQVTC QTPEQAQLPR WVAARAKQLN L ELDDAANQ VLCYCYEGNL LALAQALERL SLLWPDGKLT LPRVEQAVND AAHFTPFHWV DALLMGKSKR ALHILQQLRL EG SEPVILL RTLQRELLLL VNLKRQSAHT PLRALFDKHR VWQNRRGMMG EALNRLSQTQ LRQAVQLLTR TELTLKQDYG QSV WAELEG LSLLLCHKPL ADVFIDG UniProtKB: DNA polymerase III subunit delta |
-Macromolecule #2: DNA polymerase III subunit tau
| Macromolecule | Name: DNA polymerase III subunit tau / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO / EC number: DNA-directed DNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 47.601766 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MSYQVLARKW RPQTFADVVG QEHVLTALAN GLSLGRIHHA YLFSGTRGVG KTSIARLLAK GLNCETGITA TPCGVCDNCR EIEQGRFVD LIEIDAASRT KVEDTRDLLD NVQYAPARGR FKVYLIDEVH MLSRHSFNAL LKTLEEPPEH VKFLLATTDP Q KLPVTILS ...String: MSYQVLARKW RPQTFADVVG QEHVLTALAN GLSLGRIHHA YLFSGTRGVG KTSIARLLAK GLNCETGITA TPCGVCDNCR EIEQGRFVD LIEIDAASRT KVEDTRDLLD NVQYAPARGR FKVYLIDEVH MLSRHSFNAL LKTLEEPPEH VKFLLATTDP Q KLPVTILS RCLQFHLKAL DVEQIRHQLE HILNEEHIAH EPRALQLLAR AAEGSLRDAL SLTDQAIASG DGQVSTQAVS AM LGTLDDD QALSLVEAMV EANGERVMAL INEAAARGIE WEALLVEMLG LLHRIAMVQL SPAALGNDMA AIELRMRELA RTI PPTDIQ LYYQTLLIGR KELPYAPDRR MGVEMTLLRA LAFHPRMPLP EPEVPRQSFA PVAPTAVMTP TQVPPQPQSA PQQA PTVPL PETTSQVLAA RQQLQRVQGA TKAKKE UniProtKB: DNA polymerase III subunit tau |
-Macromolecule #3: DNA polymerase III subunit delta'
| Macromolecule | Name: DNA polymerase III subunit delta' / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed DNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.980484 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRWYPWLRPD FEKLVASYQA GRGHHALLIQ ALPGMGDDAL IYALSRYLLC QQPQGHKSCG HCRGCQLMQA GTHPDYYTLA PEKGKNTLG VDAVREVTEK LNEHARLGGA KVVWVTDAAL LTDAAANALL KTLEEPPAET WFFLATREPE RLLATLRSRC R LHYLAPPP ...String: MRWYPWLRPD FEKLVASYQA GRGHHALLIQ ALPGMGDDAL IYALSRYLLC QQPQGHKSCG HCRGCQLMQA GTHPDYYTLA PEKGKNTLG VDAVREVTEK LNEHARLGGA KVVWVTDAAL LTDAAANALL KTLEEPPAET WFFLATREPE RLLATLRSRC R LHYLAPPP EQYAVTWLSR EVTMSQDALL AALRLSAGSP GAALALFQGD NWQARETLCQ ALAYSVPSGD WYSLLAALNH EQ APARLHW LATLLMDALK RHHGAAQVTN VDVPGLVAEL ANHLSPSRLQ AILGDVCHIR EQLMSVTGIN RELLITDLLL RIE HYLQPG VVLPVPHL UniProtKB: DNA polymerase III subunit delta' |
-Macromolecule #4: DNA polymerase III subunit psi
| Macromolecule | Name: DNA polymerase III subunit psi / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO / EC number: DNA-directed DNA polymerase |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 15.188276 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTSRRDWQLQ QLGITQWSLR RPGALQGEIA IAIPAHVRLV MVANDLPALT DPLVSDVLRA LTVSPDQVLQ LTPEKIAMLP QGSHCNSWR LGTDEPLSLE GAQVASPALT DLRANPTARA ALWQQICTYE HDFFPRND UniProtKB: DNA polymerase III subunit psi |
-Macromolecule #5: Beta sliding clamp
| Macromolecule | Name: Beta sliding clamp / type: protein_or_peptide / ID: 5 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 40.630508 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MKFTVEREHL LKPLQQVSGP LGGRPTLPIL GNLLLQVADG TLSLTGTDLE MEMVARVALV QPHEPGATTV PARKFFDICR GLPEGAEIA VQLEGERMLV RSGRSRFSLS TLPAADFPNL DDWQSEVEFT LPQATMKRLI EATQFSMAHQ DVRYYLNGML F ETEGEELR ...String: MKFTVEREHL LKPLQQVSGP LGGRPTLPIL GNLLLQVADG TLSLTGTDLE MEMVARVALV QPHEPGATTV PARKFFDICR GLPEGAEIA VQLEGERMLV RSGRSRFSLS TLPAADFPNL DDWQSEVEFT LPQATMKRLI EATQFSMAHQ DVRYYLNGML F ETEGEELR TVATDGHRLA VCSMPIGQSL PSHSVIVPRK GVIELMRMLD GGDNPLRVQI GSNNIRAHVG DFIFTSKLVD GR FPDYRRV LPKNPDKHLE AGCDLLKQAF ARAAILSNEK FRGVRLYVSE NQLKITANNP EQEEAEEILD VTYSGAEMEI GFN VSYVLD VLNALKCENV RMMLTDSVSS VQIEDAASQS AAYVVMPMRL UniProtKB: Beta sliding clamp |
-Macromolecule #6: PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER
| Macromolecule | Name: PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER / type: ligand / ID: 6 / Number of copies: 3 / Formula: AGS |
|---|---|
| Molecular weight | Theoretical: 523.247 Da |
| Chemical component information |  ChemComp-AGS: |
-Macromolecule #7: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 7 / Number of copies: 3 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Macromolecule #8: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 8 / Number of copies: 4 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 Component:
Details: 30 mM Na.HEPES pH 7.5, 3 mM MgCl2, 2 mM dithiothreitol, 0.25 mM EDTA, 2% glycerol, 1 mM ATPgammaS. | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: UltrAuFoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Support film - Material: GOLD / Pretreatment - Type: PLASMA CLEANING / Pretreatment - Time: 120 sec. / Pretreatment - Atmosphere: AIR / Pretreatment - Pressure: 0.03 kPa Details: Used a Zepto Low-pressure plasma systems (PLASMA CLEANER) (Diener Electronic) at 10% power for 120 seconds. | |||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 279 K / Instrument: FEI VITROBOT MARK IV | |||||||||||||||||||||
| Details | 30 microL of 6 microM delta/tau3/delta' mixed with chi/psi complex at a molar ratio of 1:1.2, beta2 at 1:1.3, and dialysed twice at 4 degrees C against 250 mL of 30 mM Na.HEPES pH 7.5, 3 mM MgCl2, 2 mM dithiothreitol, 0.25 mM EDTA, 2% glycerol. 1 mM ATPgammaS was added to the dialysate. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Details: unfiltered mode |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Digitization - Frames/image: 1-60 / Number grids imaged: 1 / Number real images: 7269 / Average exposure time: 6.0 sec. / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 1.3 µm / Nominal defocus min: 0.4 µm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)