[English] 日本語
 Yorodumi
Yorodumi- EMDB-39226: Cryo EM structure of Komagataella phaffii RNAPII-Rat1-Rai1 pre-te... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo EM structure of Komagataella phaffii RNAPII-Rat1-Rai1 pre-termination complex | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | transcription termination / RNA polymerase II / TRANSCRIPTION | |||||||||
| Function / homology |  Function and homology information Function and homology informationmRNA 5'-diphosphatase activity / NAD-cap decapping / regulation of septum digestion after cytokinesis / co-transcriptional lncRNA 3' end processing, cleavage and polyadenylation pathway / siRNA-mediated pericentric heterochromatin formation / 5'-3' RNA exonuclease activity / nuclease activity / RPB4-RPB7 complex / intracellular phosphate ion homeostasis / chromatin-protein adaptor activity ...mRNA 5'-diphosphatase activity / NAD-cap decapping / regulation of septum digestion after cytokinesis / co-transcriptional lncRNA 3' end processing, cleavage and polyadenylation pathway / siRNA-mediated pericentric heterochromatin formation / 5'-3' RNA exonuclease activity / nuclease activity / RPB4-RPB7 complex / intracellular phosphate ion homeostasis / chromatin-protein adaptor activity / nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay / termination of RNA polymerase II transcription / termination of RNA polymerase III transcription / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / transcription initiation at RNA polymerase III promoter / termination of RNA polymerase I transcription / transcription initiation at RNA polymerase I promoter / maintenance of transcriptional fidelity during transcription elongation by RNA polymerase II / positive regulation of translational initiation / nuclear-transcribed mRNA catabolic process / RNA polymerase I complex / RNA polymerase III complex / transcription elongation by RNA polymerase I / pericentric heterochromatin / Hydrolases; Acting on acid anhydrides; In phosphorus-containing anhydrides / RNA polymerase II, core complex / tRNA transcription by RNA polymerase III / transcription by RNA polymerase I / translesion synthesis / transcription-coupled nucleotide-excision repair / translation initiation factor binding / transcription initiation at RNA polymerase II promoter / P-body / transcription elongation by RNA polymerase II / DNA-templated transcription termination / ribonucleoside binding / mRNA processing / DNA-directed RNA polymerase / rRNA processing / DNA-directed RNA polymerase activity / single-stranded DNA binding / Hydrolases; Acting on ester bonds; Exoribonucleases producing 5'-phosphomonoesters / transcription by RNA polymerase II / nucleic acid binding / protein dimerization activity / single-stranded RNA binding / nucleotide binding / RNA-directed RNA polymerase activity / DNA-templated transcription / chromatin / nucleolus / DNA binding / RNA binding / zinc ion binding / metal ion binding / nucleus / cytosol Similarity search - Function | |||||||||
| Biological species |  Komagataella phaffii (fungus) / synthetic construct (others) Komagataella phaffii (fungus) / synthetic construct (others) | |||||||||
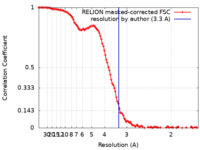
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Murayama Y / Yanagisawa T / Ehara H / Sekine S | |||||||||
| Funding support |  Japan, 1 items Japan, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural basis of eukaryotic transcription termination by the Rat1 exonuclease complex. Authors: Tatsuo Yanagisawa / Yuko Murayama / Haruhiko Ehara / Mie Goto / Mari Aoki / Shun-Ichi Sekine /  Abstract: The 5´-3´ exoribonuclease Rat1/Xrn2 is responsible for the termination of eukaryotic mRNA transcription by RNAPII. Rat1 forms a complex with its partner proteins, Rai1 and Rtt103, and acts as a ...The 5´-3´ exoribonuclease Rat1/Xrn2 is responsible for the termination of eukaryotic mRNA transcription by RNAPII. Rat1 forms a complex with its partner proteins, Rai1 and Rtt103, and acts as a "torpedo" to bind transcribing RNAPII and dissociate DNA/RNA from it. Here we report the cryo-electron microscopy structures of the Rat1-Rai1-Rtt103 complex and three Rat1-Rai1-associated RNAPII complexes (type-1, type-1b, and type-2) from the yeast, Komagataella phaffii. The Rat1-Rai1-Rtt103 structure revealed that Rat1 and Rai1 form a heterotetramer with a single Rtt103 bound between two Rai1 molecules. In the type-1 complex, Rat1-Rai1 forms a heterodimer and binds to the RNA exit site of RNAPII to extract RNA into the Rat1 exonuclease active site. This interaction changes the RNA path in favor of termination (the "pre-termination" state). The type-1b and type-2 complexes have no bound DNA/RNA, likely representing the "post-termination" states. These structures illustrate the termination mechanism of eukaryotic mRNA transcription. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_39226.map.gz emd_39226.map.gz | 116 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-39226-v30.xml emd-39226-v30.xml emd-39226.xml emd-39226.xml | 40.1 KB 40.1 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_39226_fsc.xml emd_39226_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_39226.png emd_39226.png | 77.5 KB | ||
| Filedesc metadata |  emd-39226.cif.gz emd-39226.cif.gz | 11.1 KB | ||
| Others |  emd_39226_additional_1.map.gz emd_39226_additional_1.map.gz emd_39226_half_map_1.map.gz emd_39226_half_map_1.map.gz emd_39226_half_map_2.map.gz emd_39226_half_map_2.map.gz | 28 MB 98.9 MB 98.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-39226 http://ftp.pdbj.org/pub/emdb/structures/EMD-39226 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39226 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-39226 | HTTPS FTP |
-Related structure data
| Related structure data |  8yfqMC  8yf5C  8yfeC  8yfrC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_39226.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_39226.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||
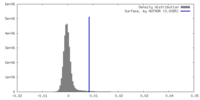
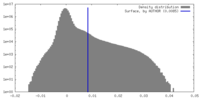
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : Cryo EM structure of Komagataella phaffii RNAPII-Rat1-Rai1 pre-te...
+Supramolecule #1: Cryo EM structure of Komagataella phaffii RNAPII-Rat1-Rai1 pre-te...
+Macromolecule #1: DNA-directed RNA polymerase subunit
+Macromolecule #2: DNA-directed RNA polymerase subunit beta
+Macromolecule #3: RNA polymerase II third largest subunit B44, part of central core
+Macromolecule #4: RNA polymerase II subunit B32
+Macromolecule #5: DNA-directed RNA polymerases I, II, and III subunit RPABC1
+Macromolecule #6: RNA polymerase subunit ABC23, common to RNA polymerases I, II, and III
+Macromolecule #7: RNA polymerase II subunit
+Macromolecule #8: DNA-directed RNA polymerases I, II, and III subunit RPABC3
+Macromolecule #9: DNA-directed RNA polymerase subunit
+Macromolecule #10: RNA polymerase subunit ABC10-beta, common to RNA polymerases I, I...
+Macromolecule #11: RNA polymerase II subunit B12.5
+Macromolecule #12: RNA polymerase subunit ABC10-alpha
+Macromolecule #16: 5'-3' exoribonuclease
+Macromolecule #17: Decapping nuclease
+Macromolecule #13: DNA (90-mer)
+Macromolecule #14: DNA (90-mer)
+Macromolecule #15: RNA (22-mer)
+Macromolecule #18: ZINC ION
+Macromolecule #19: MAGNESIUM ION
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 75 % / Chamber temperature: 283 K / Instrument: LEICA EM GP / Details: EMGP2. |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 53.3 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.6 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)